Introduction to SoilProfileCollection Objects
Source:vignettes/Introduction-to-SoilProfileCollection-Objects.Rmd
Introduction-to-SoilProfileCollection-Objects.RmdIntroduction
This is a basic introduction to the
SoilProfileCollection class object defined in the
aqp package for R.
The SoilProfileCollection class was designed to simplify
the process of working with the collection of data associated with soil
profiles: site-level data, horizon-level data, spatial data, diagnostic
horizon data, metadata, etc.
Examples listed below are meant to be copied/pasted from this
document and interactively run within R. Comments
(preceded by # symbol) briefly describe what the code in
each line does. Further documentation on objects and functions from the
aqp package can be accessed by typing
help(aqp) (or more generally, ?function_name)
at the R console.
Setup
In this tutorial we will use some sample data included with the
aqp package, based on characterization data from 10 soils
sampled on serpentinitic parent material as described in McGahan
et al, 2009.
To begin you will load required packages. You may have to first install these if missing:
# install CRAN release + dependencies
install.packages('aqp', dependencies = TRUE)
install.packages('remotes', dependencies = TRUE)
# install latest version from GitHub
remotes::install_github("ncss-tech/aqp", dependencies = FALSE)
# load sample data set, a data.frame object with horizon-level data from 10 profiles
data(sp4)
str(sp4)#> 'data.frame': 30 obs. of 13 variables:
#> $ id : chr "colusa" "colusa" "colusa" "colusa" ...
#> $ name : chr "A" "ABt" "Bt1" "Bt2" ...
#> $ top : int 0 3 8 30 0 9 0 4 13 0 ...
#> $ bottom : int 3 8 30 42 9 34 4 13 40 6 ...
#> $ K : num 0.3 0.2 0.1 0.1 0.2 0.3 0.2 0.6 0.8 0.4 ...
#> $ Mg : num 25.7 23.7 23.2 44.3 21.9 18.9 12.1 12.1 17.7 16.4 ...
#> $ Ca : num 9 5.6 1.9 0.3 4.4 4.5 1.4 7 4.4 24.1 ...
#> $ CEC_7 : num 23 21.4 23.7 43 18.8 27.5 23.7 18 20 31.1 ...
#> $ ex_Ca_to_Mg: num 0.35 0.23 0.08 0.01 0.2 0.2 0.58 0.51 0.25 1.47 ...
#> $ sand : int 46 42 40 27 54 49 43 36 27 43 ...
#> $ silt : int 33 31 28 18 20 18 55 49 45 42 ...
#> $ clay : int 21 27 32 55 25 34 3 15 27 15 ...
#> $ CF : num 0.12 0.27 0.27 0.16 0.55 0.84 0.5 0.75 0.67 0.02 ...
# optionally read about it...
# ?sp4
# upgrade to SoilProfileCollection
# 'id' is the name of the column containing the profile ID
# 'top' is the name of the column containing horizon upper boundaries
# 'bottom' is the name of the column containing horizon lower boundaries
depths(sp4) <- id ~ top + bottom
# define "horizon designation" column name for the collection
hzdesgnname(sp4) <- 'name'
# check it out:
class(sp4)#> [1] "SoilProfileCollection"
#> attr(,"package")
#> [1] "aqp"
print(sp4)#> SoilProfileCollection with 10 profiles and 30 horizons
#> profile ID: id | horizon ID: hzID
#> Depth range: 16 - 49 cm
#>
#> ----- Horizons (6 / 30 rows | 10 / 14 columns) -----
#> id hzID top bottom name K Mg Ca CEC_7 ex_Ca_to_Mg
#> colusa 1 0 3 A 0.3 25.7 9.0 23.0 0.35
#> colusa 2 3 8 ABt 0.2 23.7 5.6 21.4 0.23
#> colusa 3 8 30 Bt1 0.1 23.2 1.9 23.7 0.08
#> colusa 4 30 42 Bt2 0.1 44.3 0.3 43.0 0.01
#> glenn 5 0 9 A 0.2 21.9 4.4 18.8 0.20
#> glenn 6 9 34 Bt 0.3 18.9 4.5 27.5 0.20
#> [... more horizons ...]
#>
#> ----- Sites (6 / 10 rows | 1 / 1 columns) -----
#> id
#> colusa
#> glenn
#> kings
#> mariposa
#> mendocino
#> napa
#> [... more sites ...]
#>
#> Spatial Data:
#> [EMPTY]Object Creation
SoilProfileCollection objects are typically created by
“promoting” data.frame objects (rectangular tables of data)
that contain at least three essential columns:
- an ID column uniquely identifying groups of horizons (e.g. pedons)
- horizon top boundaries
- horizon bottom boundaries
The data.frame is sorted internally according to the
profile ID and horizon top boundary. Formula notation is used to define
the columns used to promote a data.frame object:
idcolumn ~ hz_top_column + hz_bottom_columnRapid Templating
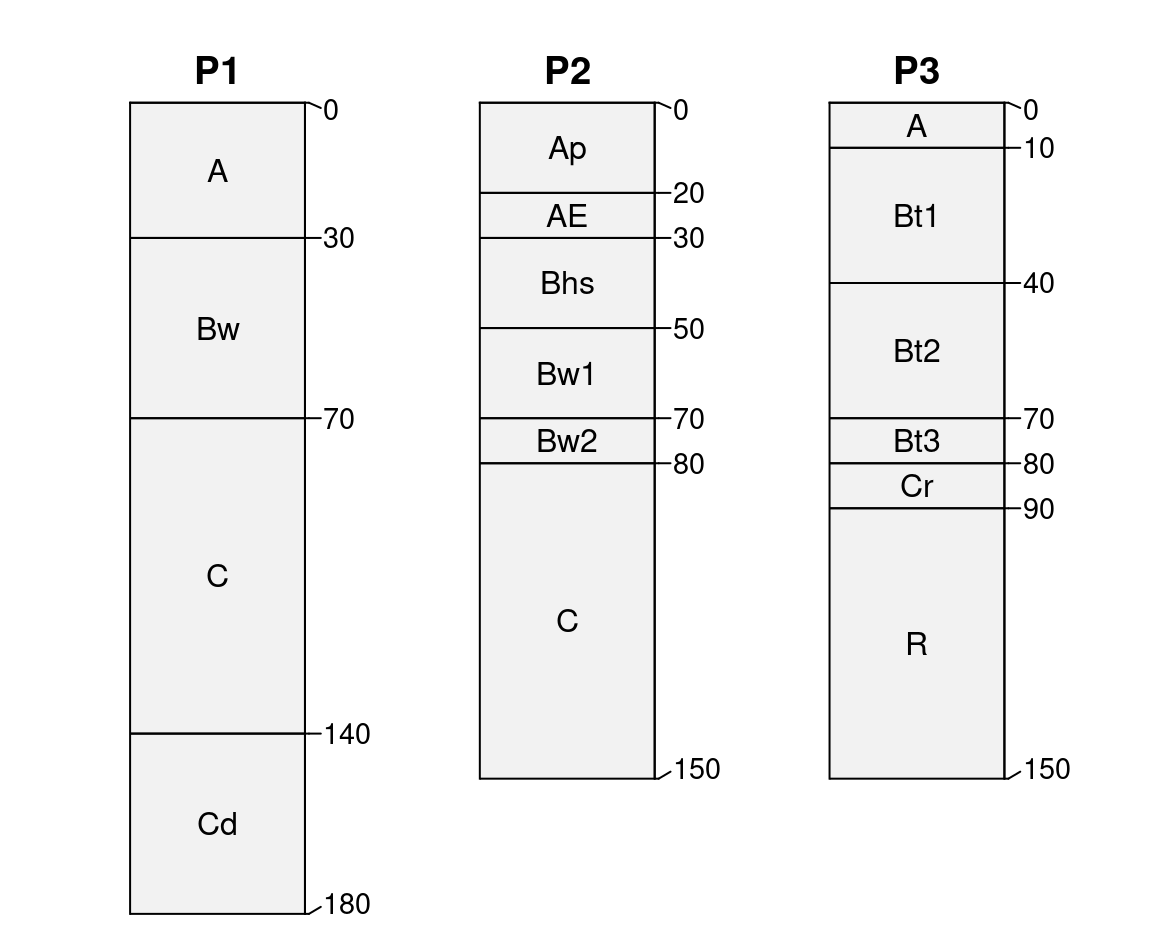
Small collections of soil profiles can be described using text
“templates” and quickSPC(). Templates take the form of:
ID:AAA|BBB|CCCCCC: relative thickness specified by horizon designations, split by “|” symbolID:A-B-C: horizons of random thickness specified by horizon designations, split by “-” symbol
The “ID:” prefix is optional, with unique IDs generated by the digest package when omitted.
# character vector of horizon templates
# must all use the same formatting
x <- c(
'P1:AAA|BwBwBwBw|CCCCCCC|CdCdCdCd',
'P2:ApAp|AE|BhsBhs|Bw1Bw1|Bw2|CCCCCCC',
'P3:A|Bt1Bt1Bt1|Bt2Bt2Bt2|Bt3|Cr|RRRRRR'
)
# each horizon label is '10' depth-units (default)
s <- quickSPC(x)
# sketch profiles
par(mar = c(0, 0, 0, 0))
plotSPC(s, name.style = 'center-center',
cex.names = 1, depth.axis = FALSE,
hz.depths = TRUE
)
Accessing, Setting, and Replacing Data
“Accessor” functions are used to extract specific components from
within SoilProfileCollection objects.
Methods that return a column name. These are useful for extracting depths, horizon designations, IDs, etc. before taking an SPC apart for a specific task.
idname(sp4): extract profile ID name (column name used to init SPC)horizonDepths(sp4): horizon top / bottom depth names (used to init SPC)hzidname(sp4): horizon ID name (typically automatically built at init time)hzdesgnname(sp4): horizon designation name (if set)hztexclname(sp4): horizon texture class name (if set)Methods that return a vector of values.
profile_id(sp4): profile IDs, in orderhzID(sp4): horizon IDs, in orderhzDesgn(sp4): horizon designations, in orderMethods that return site/horizon attribute column names.
names(sp4): site + horizon names concatenated into a single vectorhorizonNames(sp4): horizon namessiteNames(sp4): site namesProfile and horizon totals.
length(sp4): number of profiles in collectionnrow(sp4): number of horizons in collectionOther methods.
depth_units(sp4): defaults to ‘cm’ at SPC creationmetadata(sp4): returnslistobject with base + optional (user-defined) metadata elements
Horizon and Site Data
Horizon data and information about key columns (unique profile ID and depths) are the primary input to the methods provided to create the object. Site data can be derived from unique profile-specific values within the input horizon data, or joined in from an external source based on profile ID.
Both site and horizon data are stored as data.frame
within the SoilProfileCollection; with one or more rows (per profile ID)
in the horizon table and one row (per profile ID) in the site table. The
SoilProfileCollection also supports data.table, or
tibble objects in the data.frame slots.
Columns from site or horizon tables can be accessed with the
$ syntax notation, similar to the data.frame.
New data can be assigned to either table in the same manner, as long as
the length of the new data is:
- same length as the number of profiles in the collection (target is the site table)
- same length as the number of horizons in the collection (target is the horizon table)
- length 1; and selecting the target table requires
site(object)$new_column <- new_valuefor new site data andhorizons(object)$new_column <- new valuefor horizon
Assignment of new data to existing or new attributes can proceed as follows.
# site-level (based on length of assigned data == number of profiles)
sp4$elevation <- rnorm(n = length(sp4), mean = 1000, sd = 150)
# horizon-level (calculated from two horizon-level columns)
sp4$thickness <- sp4$bottom - sp4$top
# extraction of specific attributes by name
sp4$clay # vector of clay content (horizon data)#> [1] 21 27 32 55 25 34 3 15 27 32 25 31 33 13 21 23 15 17 12 19 14 14 22 25 40 51 67 24 25 32
sp4$elevation # vector of simulated elevation (site data)#> [1] 789.9935 1038.2976 634.4105 999.1643 1093.2329 1172.2617 726.7274 962.9012 963.3701
#> [10] 957.5942
# unit-length value explicitly targeting site data
site(sp4)$collection_id <- 1
# assign a single single value into horizon-level attributes
sp4$constant <- rep(1, times = nrow(sp4))
# unit-length value explicitly targeting horizon data
horizons(sp4)$analysis_group <- "SERP"
# _moves_ the named column from horizon to site
site(sp4) <- ~ constant Horizon and site data can be modified via extraction to
data.frame followed by replacement (horizon data) or join
(site data). Note that while this approach gives the most flexibility,
it is also the most dangerous–replacement of horizon data with new data
that don’t exactly conform to the original sorting may corrupt your
SoilProfileCollection.
# extract horizon data to data.frame
h <- horizons(sp4)
# add a new column and save back to original object
h$random.numbers <- rnorm(n = nrow(h), mean = 0, sd = 1)
# _replace_ original horizon data with modified version
replaceHorizons(sp4) <- h
# extract site data to data.frame
s <- site(sp4)
# add a fake group to the site data
s$group <- factor(rep(c('A', 'B'), length.out = nrow(s)))
# join new site data with previous data: old data are _not_ replaced
site(sp4) <- s
# check
sp4#> SoilProfileCollection with 10 profiles and 30 horizons
#> profile ID: id | horizon ID: hzID
#> Depth range: 16 - 49 cm
#>
#> ----- Horizons (6 / 30 rows | 10 / 17 columns) -----
#> id hzID top bottom name K Mg Ca CEC_7 ex_Ca_to_Mg
#> colusa 1 0 3 A 0.3 25.7 9.0 23.0 0.35
#> colusa 2 3 8 ABt 0.2 23.7 5.6 21.4 0.23
#> colusa 3 8 30 Bt1 0.1 23.2 1.9 23.7 0.08
#> colusa 4 30 42 Bt2 0.1 44.3 0.3 43.0 0.01
#> glenn 5 0 9 A 0.2 21.9 4.4 18.8 0.20
#> glenn 6 9 34 Bt 0.3 18.9 4.5 27.5 0.20
#> [... more horizons ...]
#>
#> ----- Sites (6 / 10 rows | 5 / 5 columns) -----
#> id elevation collection_id constant group
#> colusa 789.9935 1 1 A
#> glenn 1038.2976 1 1 B
#> kings 634.4105 1 1 A
#> mariposa 999.1643 1 1 B
#> mendocino 1093.2329 1 1 A
#> napa 1172.2617 1 1 B
#> [... more sites ...]
#>
#> Spatial Data:
#> [EMPTY]Diagnostic Horizons
Diagnostic horizons typically span several genetic horizons and may or may not be present in every profile.
To accommodate the wide range of possibilities, diagnostic horizon
data are stored as a data.frame in “long format”: each row
corresponds to a diagnostic feature in a single profile, identified with
a column matching the ID column used to initialize the
SoilProfileCollection object and a label reflecting the
feature kind.
For diagnostic horizon data there are no restrictions on data content, as long as each row has an ID that exists within the collection. Be sure to use the ID column name that was used to initialize the SoilProfileCollection object.
dh <- data.frame(
id = 'colusa',
kind = 'argillic',
top = 8,
bottom = 42,
stringsAsFactors = FALSE
)
# overwrite any existing diagnostic horizon data
diagnostic_hz(sp4) <- dh
# append to diagnostic horizon data
dh <- diagnostic_hz(sp4)
dh.new <- data.frame(
id = 'napa',
kind = 'argillic',
top = 6,
bottom = 20,
stringsAsFactors = FALSE
)
# overwrite existing diagnostic horizon data with appended data
diagnostic_hz(sp4) <- rbind(dh, dh.new)Root Restrictive Features
Features that restrict root entry (fine or very fine roots) are commonly used to estimate functional soil depth. Restrictive features include salt accumulations, duripans, fragipans, paralithic materials, lithic contact, or an abrupt change in chemical property.
Not all soils have restrictive features, therefore these data are
stored as a data.frame in “long format”. Each row
corresponds to a restrictive feature, associated depths, and identified
by profile_id(). There may be more than one restrictive
feature per soil profile.
The example data sp4 does not describe the restrictive
features, so we will simulate some at the bottom of each profile +
20cm.
# get the depth of each profile
rf.top <- profileApply(sp4, max)
rf.bottom <- rf.top + 20
# the profile IDs can be extracted from the names attribute
pIDs <- names(rf.top)
# note: profile IDs must be stored in a column named for idname(sp4) -> 'id'
rf <- data.frame(
id = pIDs,
top = rf.top,
bottom = rf.bottom,
kind = 'fake',
stringsAsFactors = FALSE
)
# overwrite any existing diagnostic horizon data
restrictions(sp4) <- rf
# check
restrictions(sp4)#> id top bottom kind
#> 1 colusa 42 62 fake
#> 2 glenn 34 54 fake
#> 3 kings 40 60 fake
#> 4 mariposa 49 69 fake
#> 5 mendocino 30 50 fake
#> 6 napa 20 40 fake
#> 7 san benito 20 40 fake
#> 8 shasta 40 60 fake
#> 9 shasta-trinity 40 60 fake
#> 10 tehama 16 36 fakeObject Metadata
SoilProfileCollection metadata can be extracted and set using the
metadata() and metadata<- methods.
#> List of 6
#> $ aqp_df_class : chr "data.frame"
#> $ aqp_group_by : chr ""
#> $ aqp_hzdesgn : chr "name"
#> $ aqp_hztexcl : chr ""
#> $ depth_units : chr "cm"
#> $ stringsAsFactors: logi FALSE
# alter the depth unit metadata attribute
depth_units(sp4) <- 'inches' # units are really 'cm'
# add or replace custom metadata
metadata(sp4)$describer <- 'DGM'
metadata(sp4)$date <- as.Date('2009-01-01')
metadata(sp4)$citation <- 'McGahan, D.G., Southard, R.J, Claassen, V.P. 2009. Plant-Available Calcium Varies Widely in Soils on Serpentinite Landscapes. Soil Sci. Soc. Am. J. 73: 2087-2095.'
# check new values have been added
str(metadata(sp4))#> List of 9
#> $ aqp_df_class : chr "data.frame"
#> $ aqp_group_by : chr ""
#> $ aqp_hzdesgn : chr "name"
#> $ aqp_hztexcl : chr ""
#> $ depth_units : chr "inches"
#> $ stringsAsFactors: logi FALSE
#> $ describer : chr "DGM"
#> $ date : Date[1:1], format: "2009-01-01"
#> $ citation : chr "McGahan, D.G., Southard, R.J, Claassen, V.P. 2009. Plant-Available Calcium Varies Widely in Soils on Serpentini"| __truncated__
# fix depth units, set back to 'cm'
depth_units(sp4) <- 'cm'Spatial Data
Spatial data and metadata can be stored within a
SoilProfileCollection object. To initialize a “spatial”
SoilProfileCollection, use initSpatial()<- method. This
allows you to specify which columns contain the geometry and optionally
the Coordinate Reference System of those data. The SoilProfileCollection
stores two metadata entries: coordinates and
crs which refer to the column names containing geometric
data and the CRS, respectively.
# generate some fake coordinates as site level attributes
sp4$x <- rnorm(n = length(sp4), mean = 354000, sd = 100)
sp4$y <- rnorm(n = length(sp4), mean = 4109533, sd = 100)
# initialize spatial coordinates (CRS optional)
initSpatial(sp4, crs = "EPSG:26911") <- ~ x + y
# extract coordinates as matrix
getSpatial(sp4)#> x y
#> [1,] 354007.0 4109619
#> [2,] 353936.1 4109509
#> [3,] 353995.0 4109512
#> [4,] 353974.9 4109535
#> [5,] 354044.5 4109536
#> [6,] 354275.5 4109588
#> [7,] 354004.7 4109306
#> [8,] 354057.8 4109801
#> [9,] 354011.8 4109497
#> [10,] 353808.8 4109554
# get/set spatial reference system using prj()<-
prj(sp4) <- '+proj=utm +zone=11 +datum=NAD83'
# return CRS information
prj(sp4)#> [1] "+proj=utm +zone=11 +datum=NAD83"
if (requireNamespace("sf", quietly = TRUE)) {
# extract spatial data + site level attributes in new spatial object
sp4.sp <- as(sp4, 'SpatialPointsDataFrame')
sp4.sf <- as(sp4, 'sf')
}Validity and Horizon Logic
There are several SoilProfileCollection methods defined to identify corrupted objects/bad depth logic and fix them.
Usually an “invalid” or “corrupt” SoilProfileCollection comes from direct modification of the S4 object slot contents (not using standard methods), which can result in omissions or reordering that bypasses internal accounting methods.
Occasionally issues arise from illogical inputs (depths) or collections that are missing data. The latter cases (standard methods produce a corrupt SPC given some “degenerate” input data) are considered “bugs” that should reported on the issue tracker: https://github.com/ncss-tech/aqp/issues
checkSPC() returns TRUE for a SoilProfileCollection that
contains all slots defined in the class prototype.
checkSPC(sp4)spc_in_sync() is used as the validity method for the
SoilProfileCollection, it determines if some reordering of the horizon
data relative to the unique profile ID / site order has occurred.
spc_in_sync(sp4)#> nSites relativeOrder valid
#> 1 TRUE TRUE TRUErebuildSPC() is used to re-construct all the required
slots of a SoilProfileCollection, given a source, possibly corrupt,
object. This function fixes major issues related to the internal
ordering of data in slots, as well as missing slots or metadata.
z <- rebuildSPC(sp4)checkHzDepthLogic() has the ability to perform logical
tests on whole profiles or individual horizons. Four different tests are
performed related to four common errors in horizon depths:
bottom depth shallower than top depth
equal top and bottom depth
missing (
NA) top or bottom depthgap or overlap between adjacent horizons
checkHzDepthLogic(sp4)#> id valid depthLogic sameDepth missingDepth overlapOrGap
#> 1 colusa TRUE FALSE FALSE FALSE FALSE
#> 2 glenn TRUE FALSE FALSE FALSE FALSE
#> 3 kings TRUE FALSE FALSE FALSE FALSE
#> 4 mariposa TRUE FALSE FALSE FALSE FALSE
#> 5 mendocino TRUE FALSE FALSE FALSE FALSE
#> 6 napa TRUE FALSE FALSE FALSE FALSE
#> 7 san benito TRUE FALSE FALSE FALSE FALSE
#> 8 shasta TRUE FALSE FALSE FALSE FALSE
#> 9 shasta-trinity TRUE FALSE FALSE FALSE FALSE
#> 10 tehama TRUE FALSE FALSE FALSE FALSE
checkHzDepthLogic(sp4, byhz = TRUE)#> id top bottom valid hzID depthLogic sameDepth missingDepth overlapOrGap
#> 1 colusa 0 3 TRUE 1 FALSE FALSE FALSE NA
#> 2 colusa 3 8 TRUE 2 FALSE FALSE FALSE NA
#> 3 colusa 8 30 TRUE 3 FALSE FALSE FALSE NA
#> 4 colusa 30 42 TRUE 4 FALSE FALSE FALSE NA
#> 5 glenn 0 9 TRUE 5 FALSE FALSE FALSE NA
#> 6 glenn 9 34 TRUE 6 FALSE FALSE FALSE NA
#> 7 kings 0 4 TRUE 7 FALSE FALSE FALSE NA
#> 8 kings 4 13 TRUE 8 FALSE FALSE FALSE NA
#> 9 kings 13 40 TRUE 9 FALSE FALSE FALSE NA
#> 10 mariposa 0 3 TRUE 10 FALSE FALSE FALSE NA
#> 11 mariposa 3 14 TRUE 11 FALSE FALSE FALSE NA
#> 12 mariposa 14 34 TRUE 12 FALSE FALSE FALSE NA
#> 13 mariposa 34 49 TRUE 13 FALSE FALSE FALSE NA
#> 14 mendocino 0 2 TRUE 14 FALSE FALSE FALSE NA
#> 15 mendocino 2 8 TRUE 15 FALSE FALSE FALSE NA
#> 16 mendocino 8 30 TRUE 16 FALSE FALSE FALSE NA
#> 17 napa 0 6 TRUE 17 FALSE FALSE FALSE NA
#> 18 napa 6 20 TRUE 18 FALSE FALSE FALSE NA
#> 19 san benito 0 8 TRUE 19 FALSE FALSE FALSE NA
#> 20 san benito 8 20 TRUE 20 FALSE FALSE FALSE NA
#> 21 shasta 0 3 TRUE 21 FALSE FALSE FALSE NA
#> 22 shasta 3 40 TRUE 22 FALSE FALSE FALSE NA
#> 23 shasta-trinity 0 2 TRUE 23 FALSE FALSE FALSE NA
#> 24 shasta-trinity 2 5 TRUE 24 FALSE FALSE FALSE NA
#> 25 shasta-trinity 5 12 TRUE 25 FALSE FALSE FALSE NA
#> 26 shasta-trinity 12 23 TRUE 26 FALSE FALSE FALSE NA
#> 27 shasta-trinity 23 40 TRUE 27 FALSE FALSE FALSE NA
#> 28 tehama 0 3 TRUE 28 FALSE FALSE FALSE NA
#> 29 tehama 3 7 TRUE 29 FALSE FALSE FALSE NA
#> 30 tehama 7 16 TRUE 30 FALSE FALSE FALSE NACoercion
SoilProfileCollection objects can be coerced to
data.frame, list and
SpatialPointsDataFrame (when spatial slot has been set
up):
# check our work by viewing the internal structure
str(sp4)
# create a data.frame from horizon+site data
as(sp4, 'data.frame')
# or, equivalently:
as.data.frame(sp4)
# convert SoilProfileCollection to a named list containing all slots
as(sp4, 'list')
# extraction of site + spatial data as SpatialPointsDataFrame
as(sp4, 'SpatialPointsDataFrame')Subsetting SoilProfileCollection Objects
SoilProfileCollection objects can be subset using the
familiar [-style notation used by matrix and
data.frame objects, such that: spc[i, j] will
return profiles identified by the integer vector i, and
horizons identified by the integer vector j.
Omitting either index will result in all profiles (i
omitted) or all horizons (j omitted).
Typically, site-level attributes will be used as the subsetting
criteria. Functions that return an index to matches (such as
grep() or which()) provide the link between
attributes and an index to matching profiles.
Profiles
Using the i index, select one or more profiles by numeric
index. An index greater than the number of profiles will return an empty
SoilProfileCollection object.
# explicit string matching
idx <- which(sp4$group == 'A')
# numerical expressions
idx <- which(sp4$elevation < 1000)
# regular expression, matches any profile ID containing 'shasta'
idx <- grep('shasta', profile_id(sp4), ignore.case = TRUE)
# perform subset based on index
sp4[idx, ]In an interactive session, it is often simpler to use
subset() directly:
Horizons by Index
Using the j index, select the 2nd horizon from each profile in the collection. Use with caution, asking for a horizon beyond the length of any profile in the collection will result in an error.
sp4[, 2]Horizons by Keyword
Using the k index, select the .FIRST first or
.LAST horizons (by profile) within a collection. Note that
.FIRST and .LAST are special keywords used by
the SoilProfileCollection subset methods.
sp4[, , .FIRST]
sp4[, , .LAST]Additional k index keywords include: .HZID,
.NHZ, .BOTTOM, .TOP. These can be
chained together to get the “top depth of the first horizons” or
“row-index (horizon data) of the last horizons”:
sp4[, , .FIRST, .TOP]
sp4[, , .LAST, .HZID]Splitting, Duplication, and Selection of Unique Profiles
SoilProfileCollection objects are combined by passing a
list of objects to the combine() function.
Ideally all objects share the same internal structure, profile ID,
horizon ID, depth units, and other parameters of a
SoilProfileCollection. Manually subset the example data
into 3 pieces, compile into a list, and then combine back
together.
# subset data into chunks
s1 <- sp4[1:2, ]
s2 <- sp4[4, ]
s3 <- sp4[c(6, 8, 9), ]
# combine subsets
s <- combine(list(s1, s2, s3))
# double-check result
plotSPC(s)It is possible to combine SoilProfileCollection objects
with different internal structure. The final object will contain the all
site and horizon columns from the inputs, possibly creating sparse
tables. IDs and horizon depth names are taken from the first object.
# sample data as data.frame objects
data(sp1)
data(sp3)
# rename IDs horizon top / bottom columns
sp3$newid <- sp3$id
sp3$hztop <- sp3$top
sp3$hzbottom <- sp3$bottom
# remove originals
sp3$id <- NULL
sp3$top <- NULL
sp3$bottom <- NULL
# promote to SoilProfileCollection
depths(sp1) <- id ~ top + bottom
depths(sp3) <- newid ~ hztop + hzbottom
# label each group via site-level attribute
site(sp1)$g <- 'sp1'
site(sp3)$g <- 'sp3'
# combine
x <- combine(list(sp1, sp3))
# make grouping variable into a factor for groupedProfilePlot
x$g <- factor(x$g)
# check results
str(x)
# graphical check
# convert character horizon IDs into numeric
x$.horizon_ids_numeric <- as.numeric(hzID(x))
par(mar = c(0, 0, 3, 1))
plotSPC(x, color='.horizon_ids_numeric', col.label = 'Horizon ID')
groupedProfilePlot(x, 'g', color='.horizon_ids_numeric', col.label = 'Horizon ID', group.name.offset = -15)Splitting
The inverse of combine() is split():
subsets of the SoilProfileCollection are split into list
elements, each containing a new SoilProfileCollection.
Duplication
Duplicate the first profile in sp4 (Colusa) 8 times,
resulting in a new SoilProfileCollection object containing
unique profile IDs.
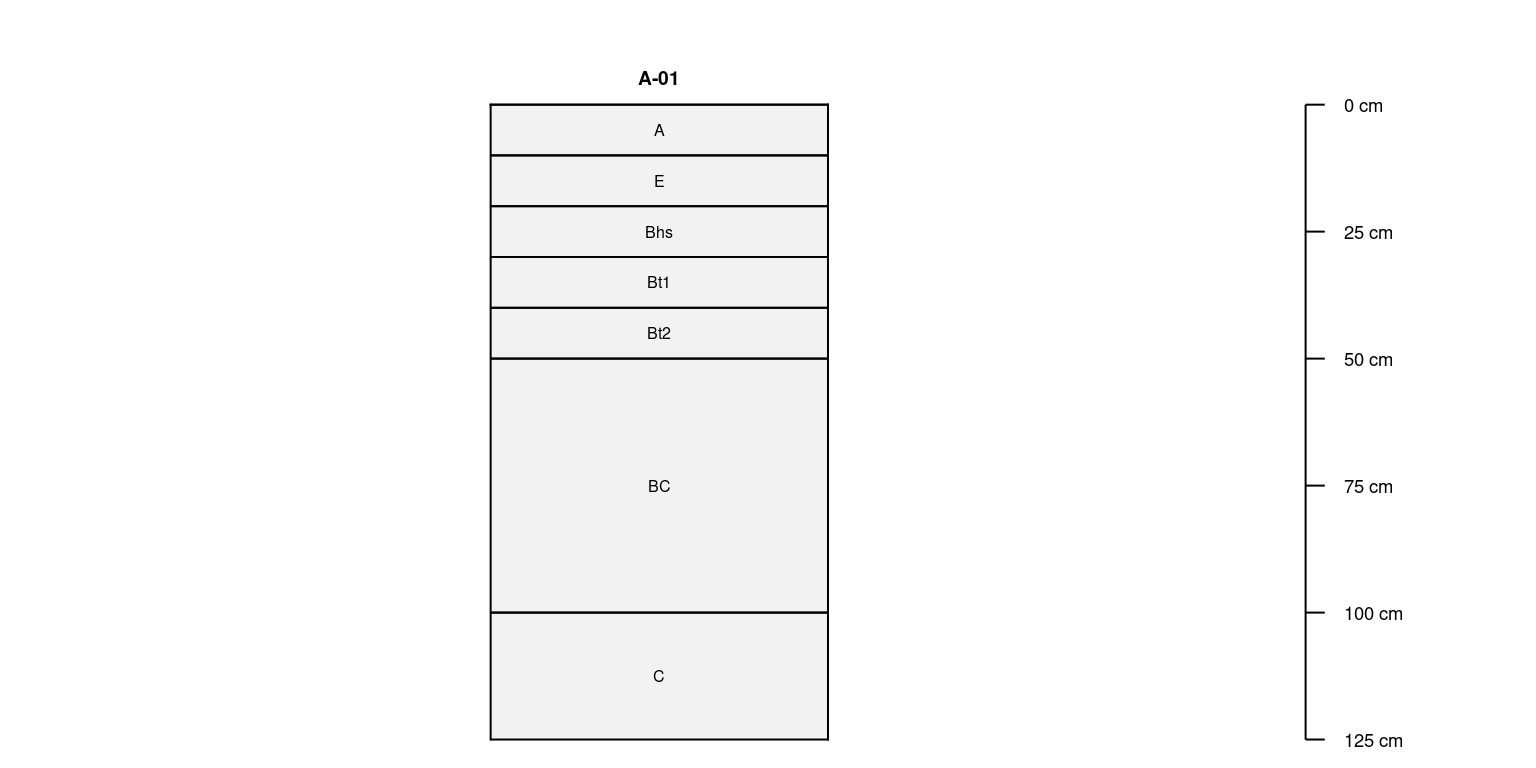

Selecting Unique Profiles
# an example soil profile
x <- data.frame(
id = 'A',
name = c('A', 'E', 'Bhs', 'Bt1', 'Bt2', 'BC', 'C'),
top = c(0, 10, 20, 30, 40, 50, 100),
bottom = c(10, 20, 30, 40, 50, 100, 125),
z = c(8, 5, 3, 7, 10, 2, 12)
)
# init SPC
depths(x) <- id ~ top + bottom
hzdesgnname(x) <- 'name'
# horizon depth variability for simulation
horizons(x)$.sd <- 2
# duplicate several times
x.dupes <- duplicate(x, times = 5)
# simulate some new profiles based on example
# 2cm constant standard deviation of transition between horizons assumed
x.sim <- perturb(x, n = 5, thickness.attr = '.sd')
# graphical check
par(mar = c(0, 2, 0, 4))
# inspect unique results
plotSPC(unique(x.dupes, vars = c('top', 'bottom')),
name.style = 'center-center',
width = 0.15)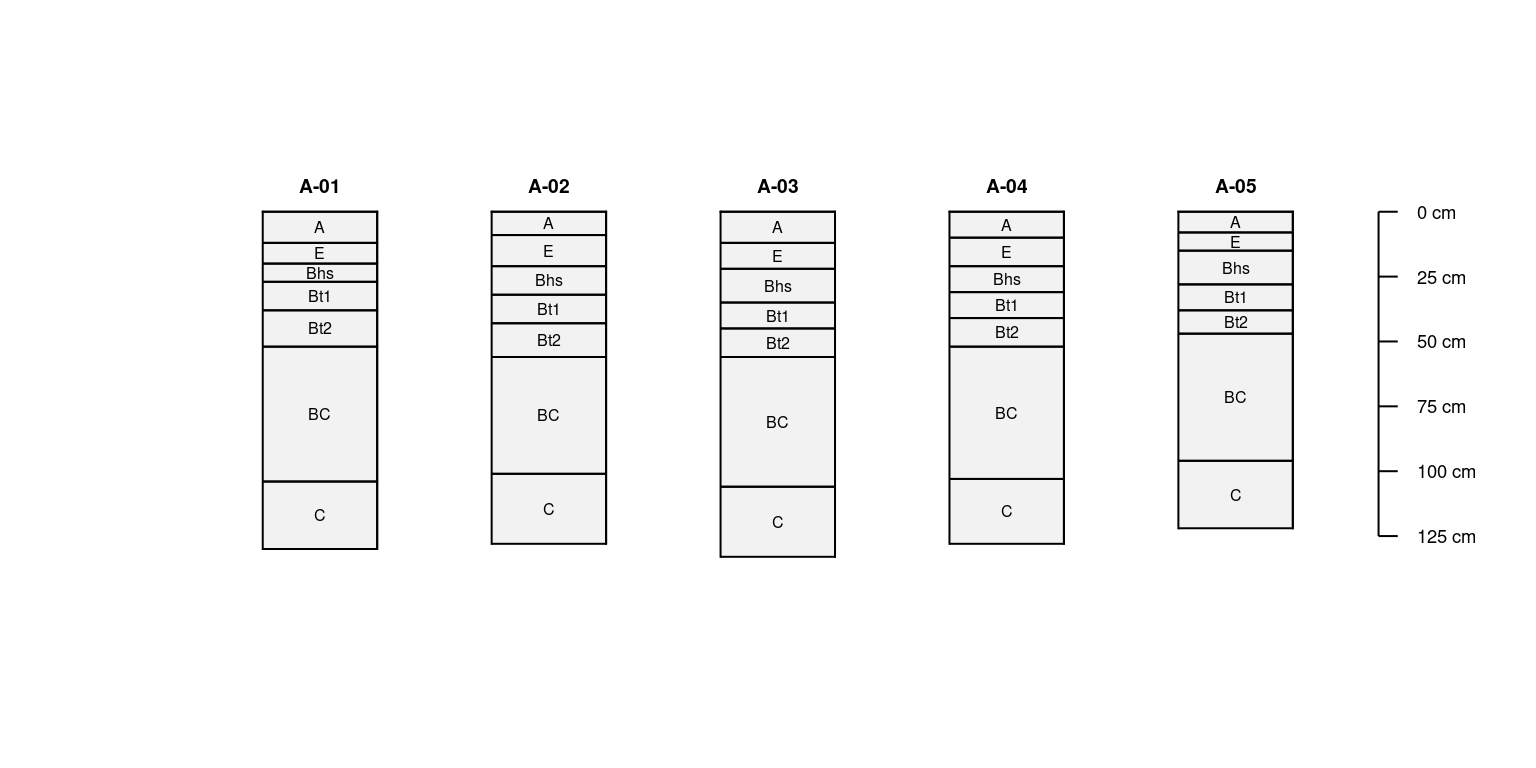
# "uniqueness" is a function of variables selected to consider
plotSPC(unique(x.sim, vars = c('top', 'bottom')),
name.style = 'center-center')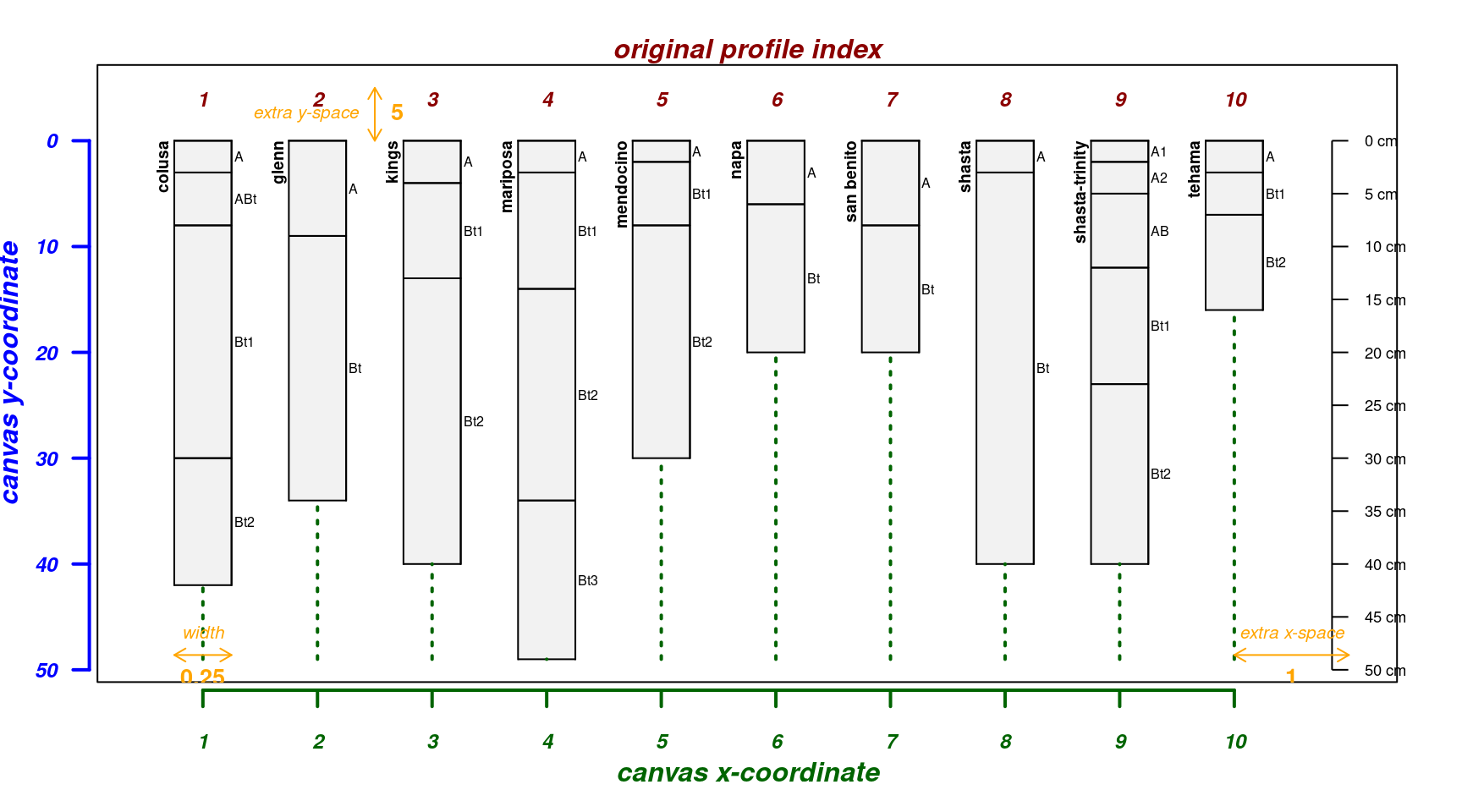

Plotting SoilProfileCollection Objects
The plotSPC() method for
SoilProfileCollection objects generates sketches of
profiles within the collection based on horizon boundaries, vertically
aligned to an integer sequence from 1 to the number of profiles.
Horizon names are automatically extracted from a horizon-level
attribute name (if present), or via an alternate attributed
given as an argument: name='column.name'.
Horizon colors are automatically generated from the horizon-level
attribute soil_color, or any other attribute of
R-compatible color description given as an argument:
color='column.name'. This function is highly customizable,
therefore, it is prudent to consult help(plotSPC) from time
to time. Soil colors in Munsell notation can be converted to
R-compatible colors via munsell2rgb().
Many of the following examples make use of par() to set
graphical options such as mar (customized margins) and/or
xpd = NA (turn off clipping) to optimize display of
different numbers of profiles and various plotSPC()
arguments.
A detailed description of each argument to plotSPC()
(and many examples) can be found in the Soil Profile
Sketches tutorial.
Making Adjustments
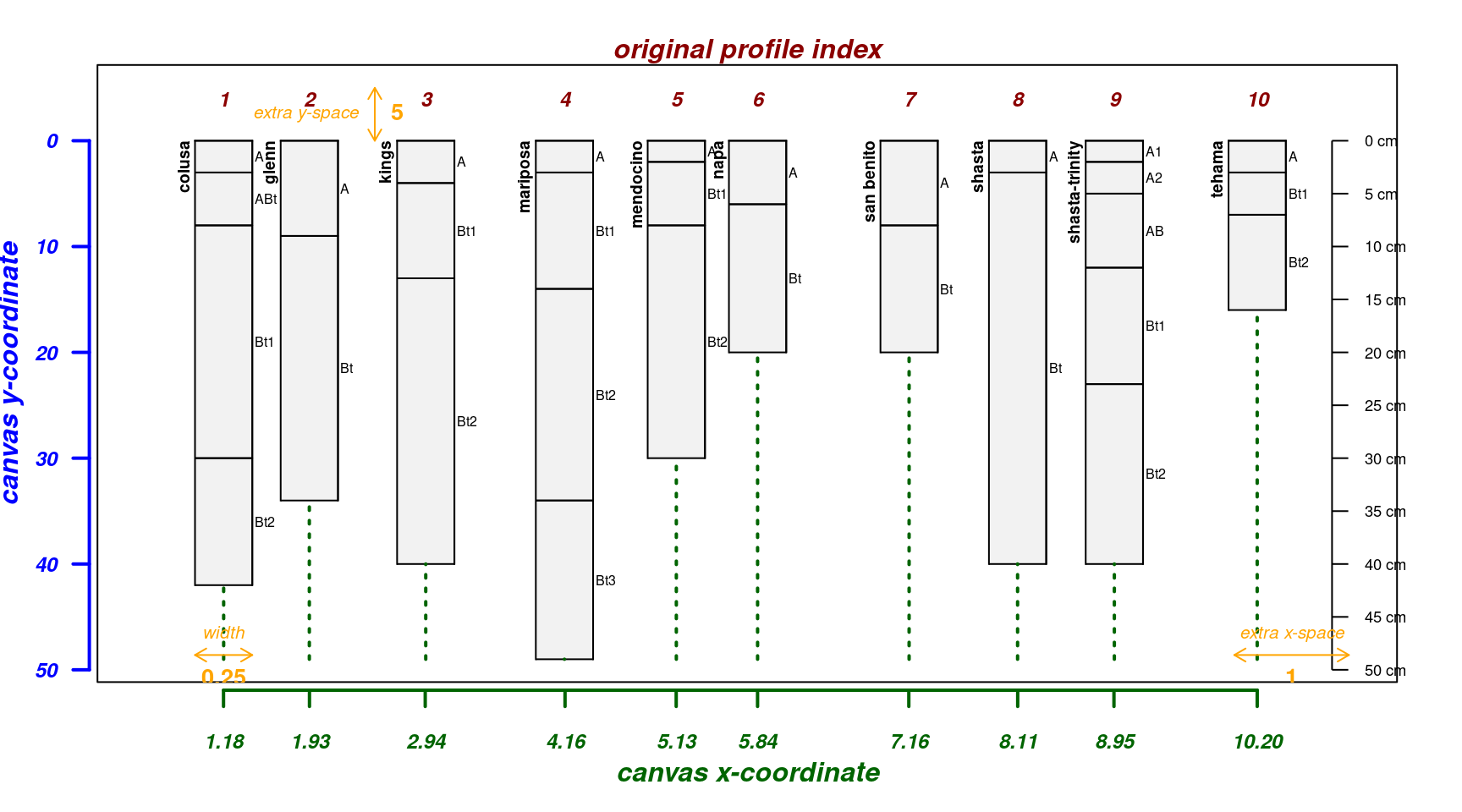
The explainPlotSPC() function is helpful for adjusting
some of the more obscure arguments to plotSPC().
A basic plot with debugging information overlaid:
par(mar = c(4, 3, 2, 2))
explainPlotSPC(sp4, name = 'name')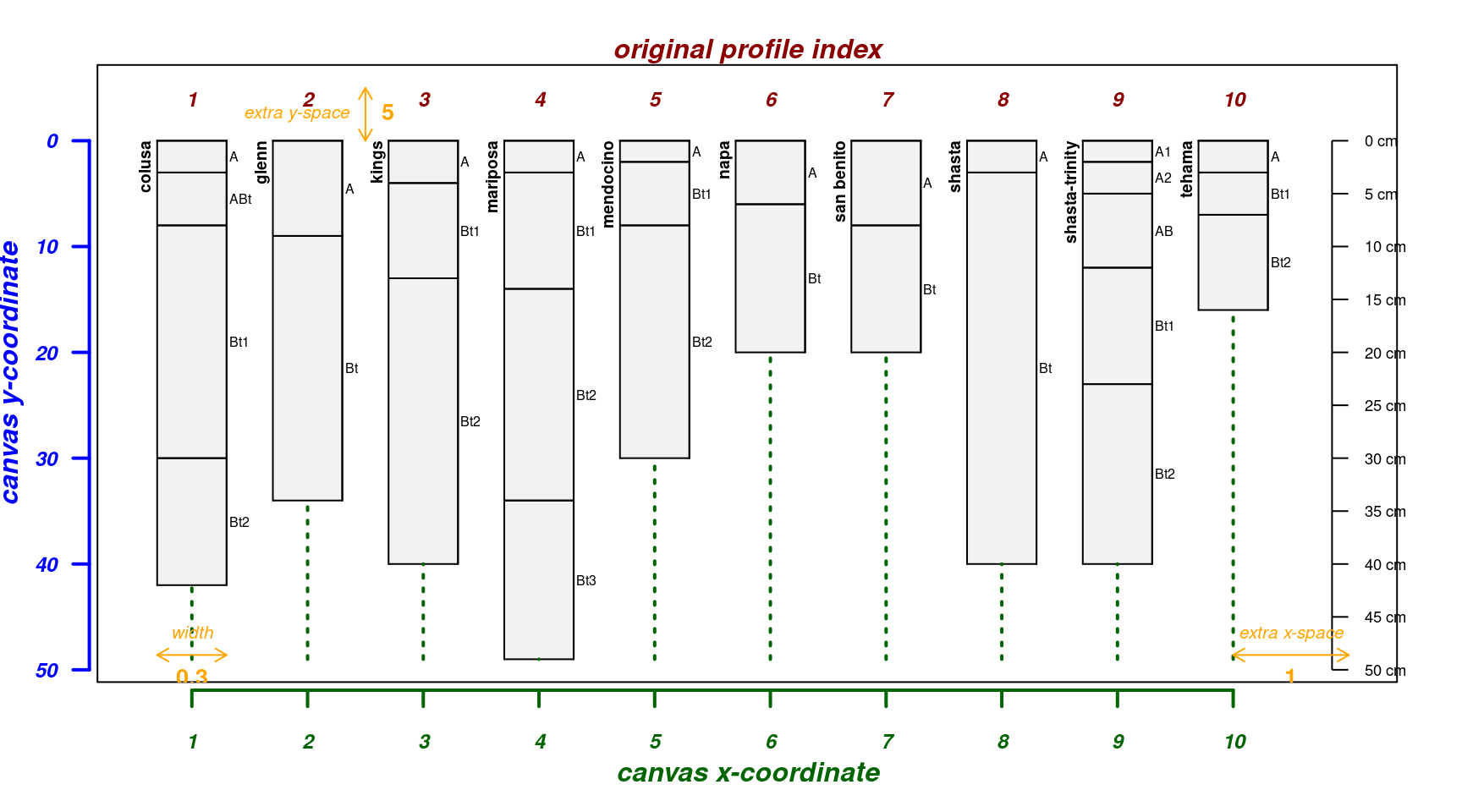
Make sketches wider:
par(mar = c(4, 3, 2, 2))
explainPlotSPC(sp4, name = 'name', width = 0.3)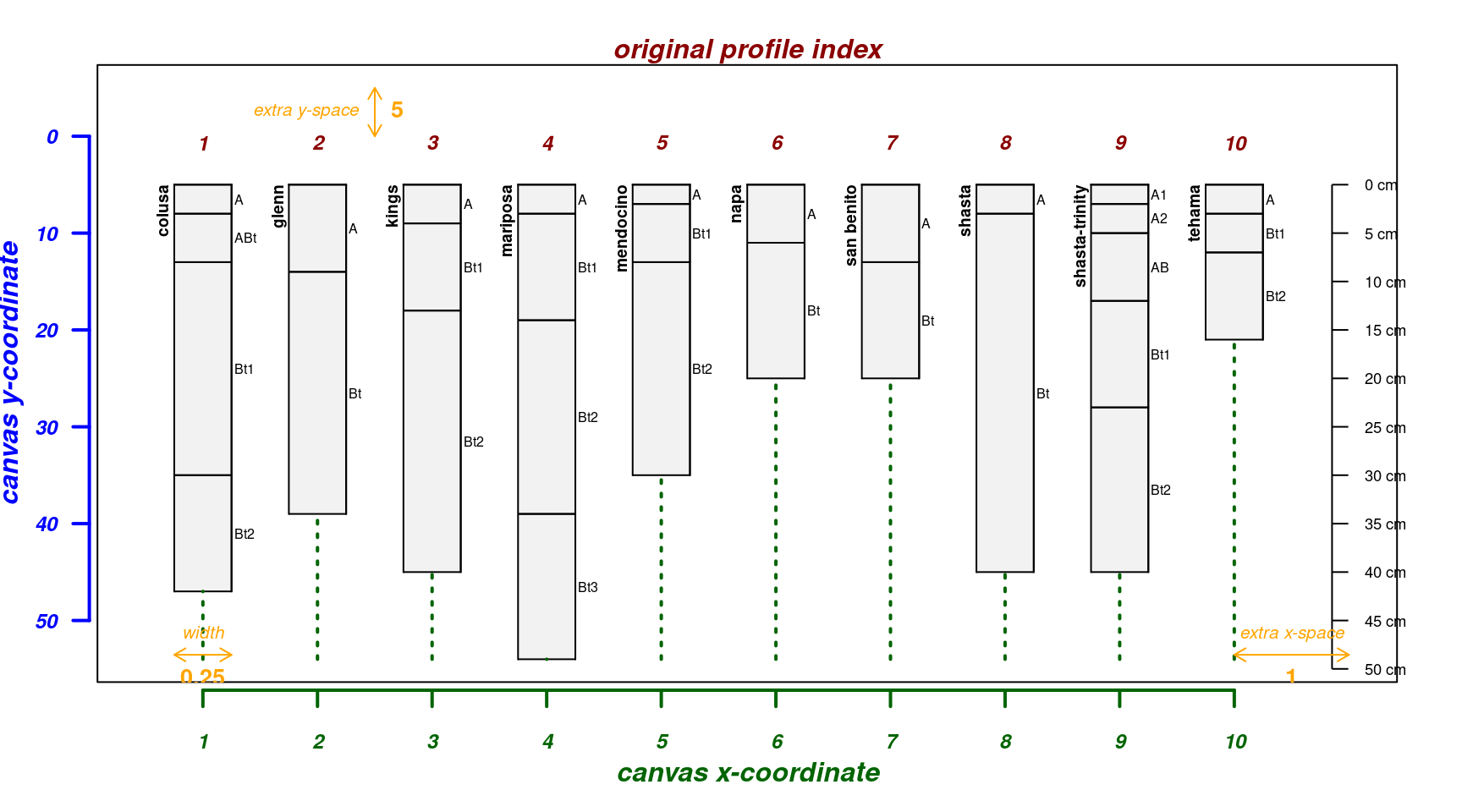
Move soil surface at 0cm “down” 5cm:
par(mar = c(4, 3, 2, 2))
explainPlotSPC(sp4, name = 'name', y.offset = 5)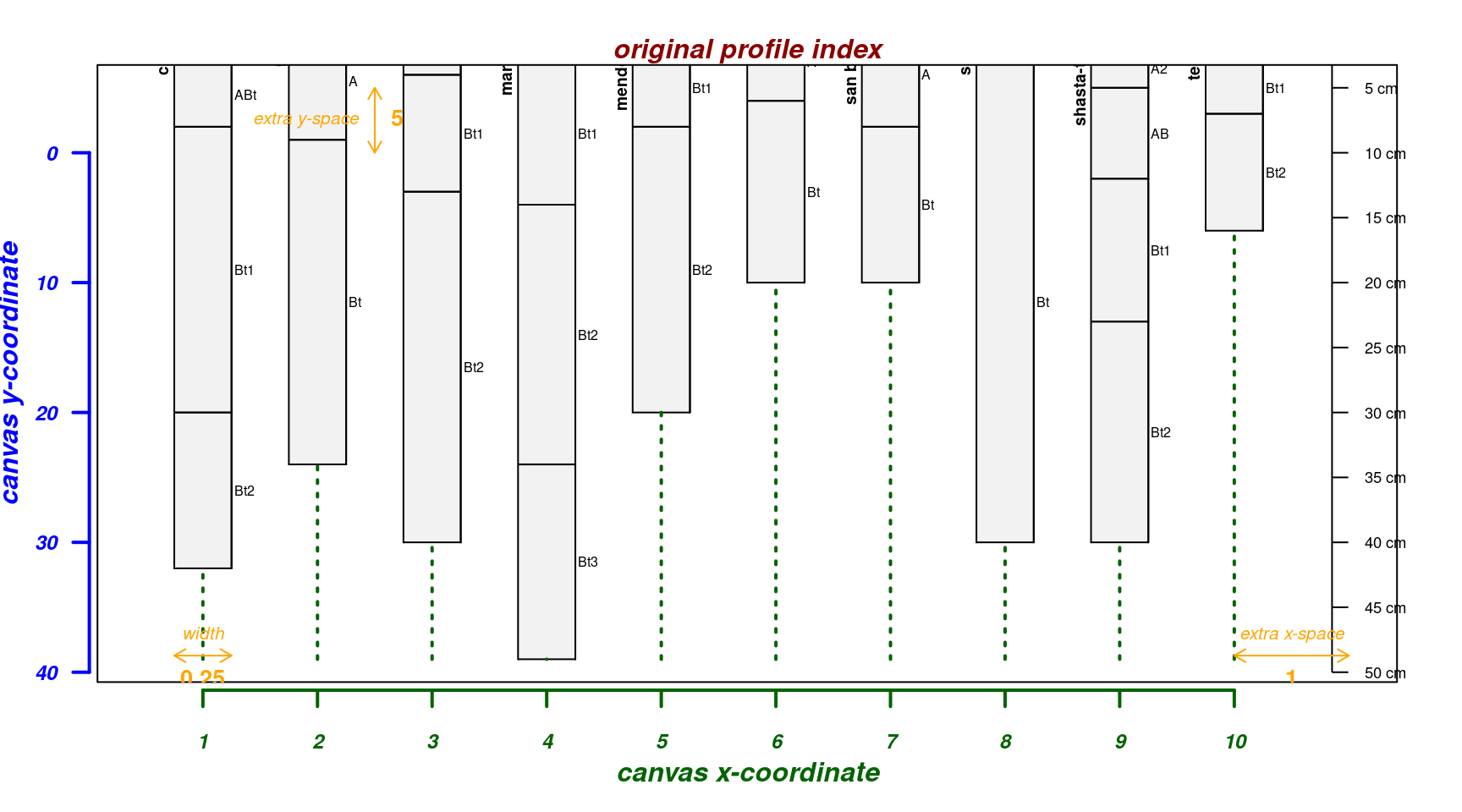
Move soil surface at 0cm “up” 10cm; useful for sketches of shallow profiles:
par(mar = c(4, 3, 2, 2))
explainPlotSPC(sp4, name = 'name', y.offset = -10)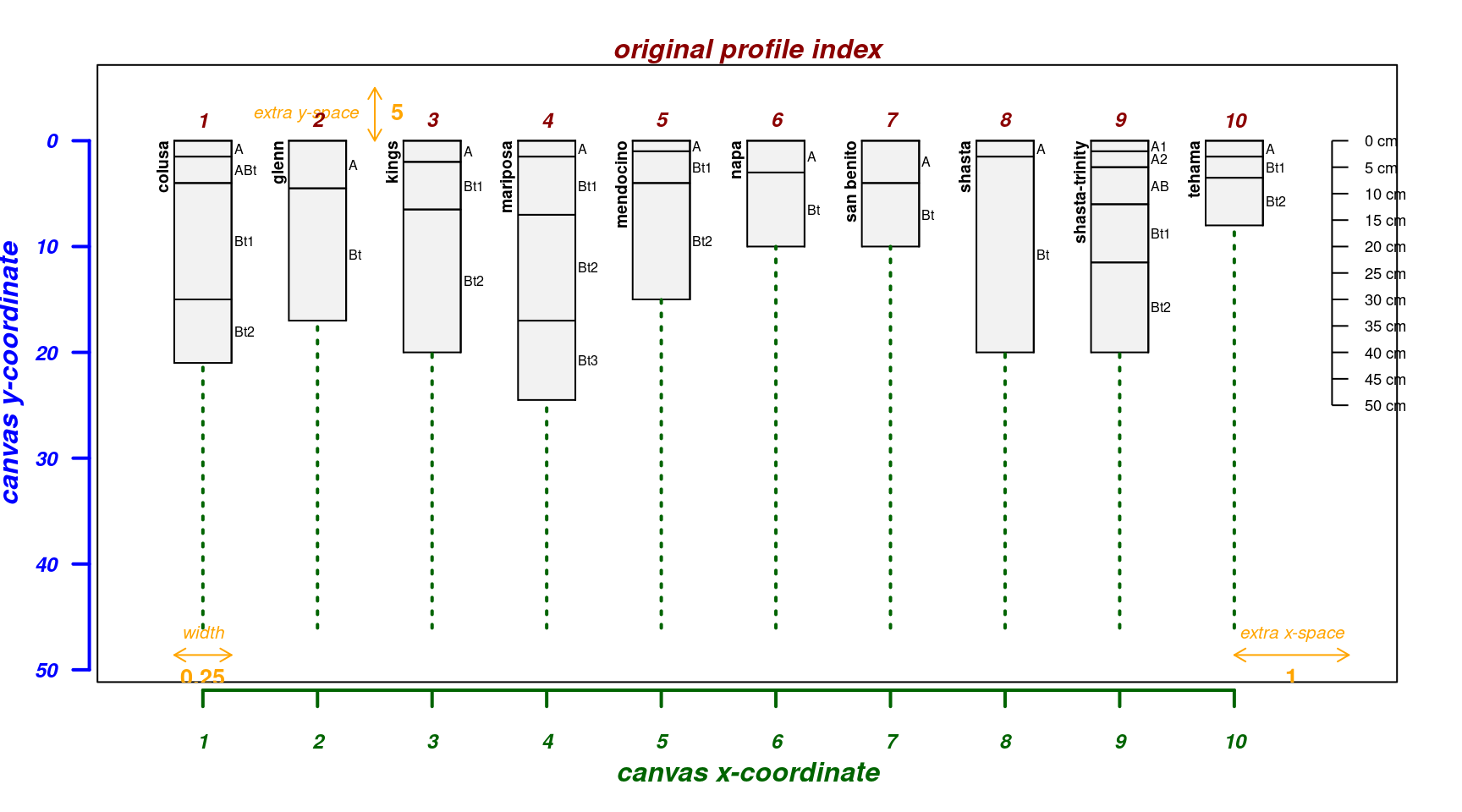
Scale depths by 50%:
par(mar = c(4, 3, 2, 2))
explainPlotSPC(sp4, name = 'name', scaling.factor = 0.5)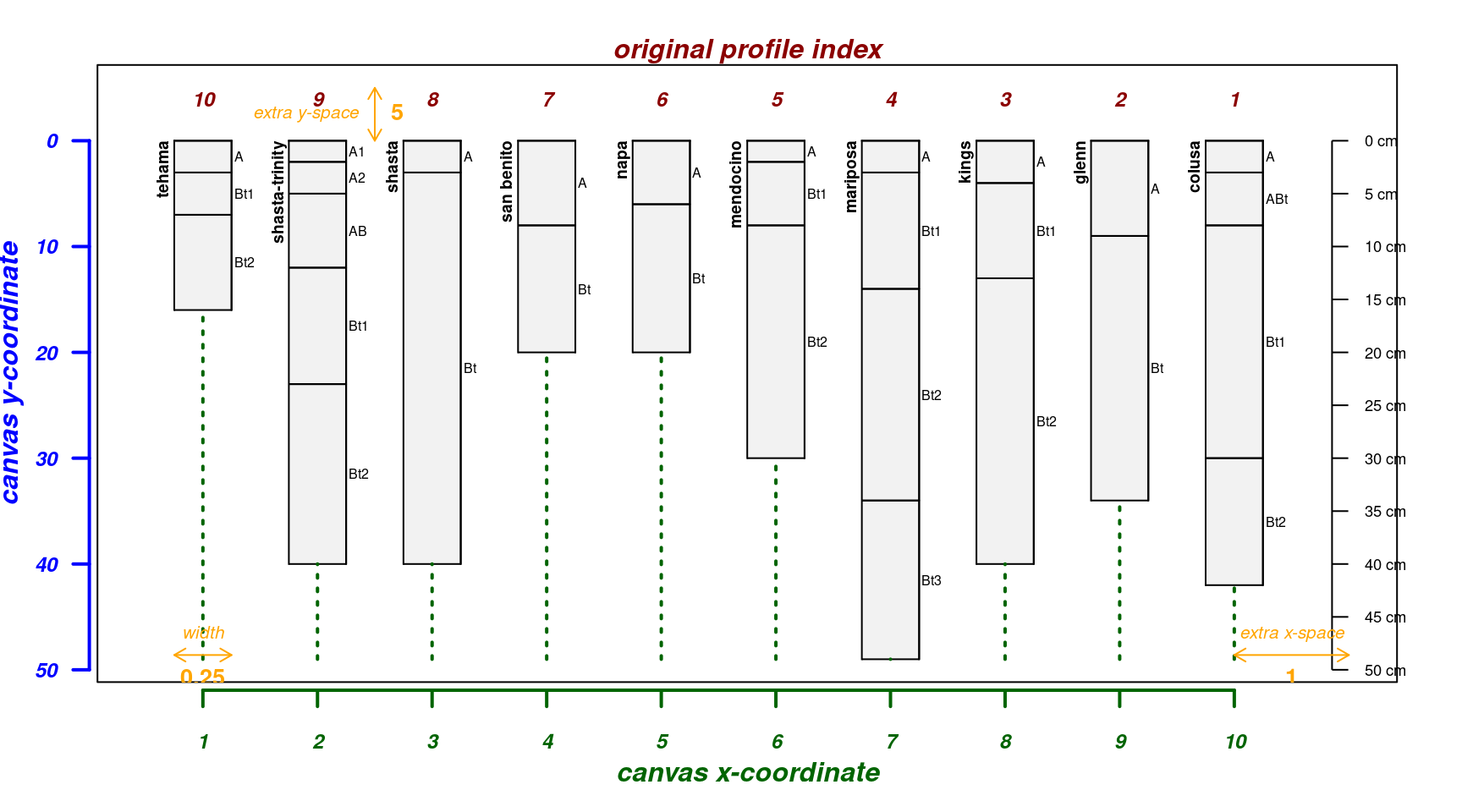
A graphical explanation of how profiles are re-arranged via
plot.order argument:
par(mar = c(4, 3, 2, 2))
explainPlotSPC(sp4, name = 'name', plot.order = length(sp4):1)
Leave room for an additional 2 profile sketches:
par(mar = c(4, 3, 2, 2))
explainPlotSPC(sp4, name = 'name', n = length(sp4) + 2)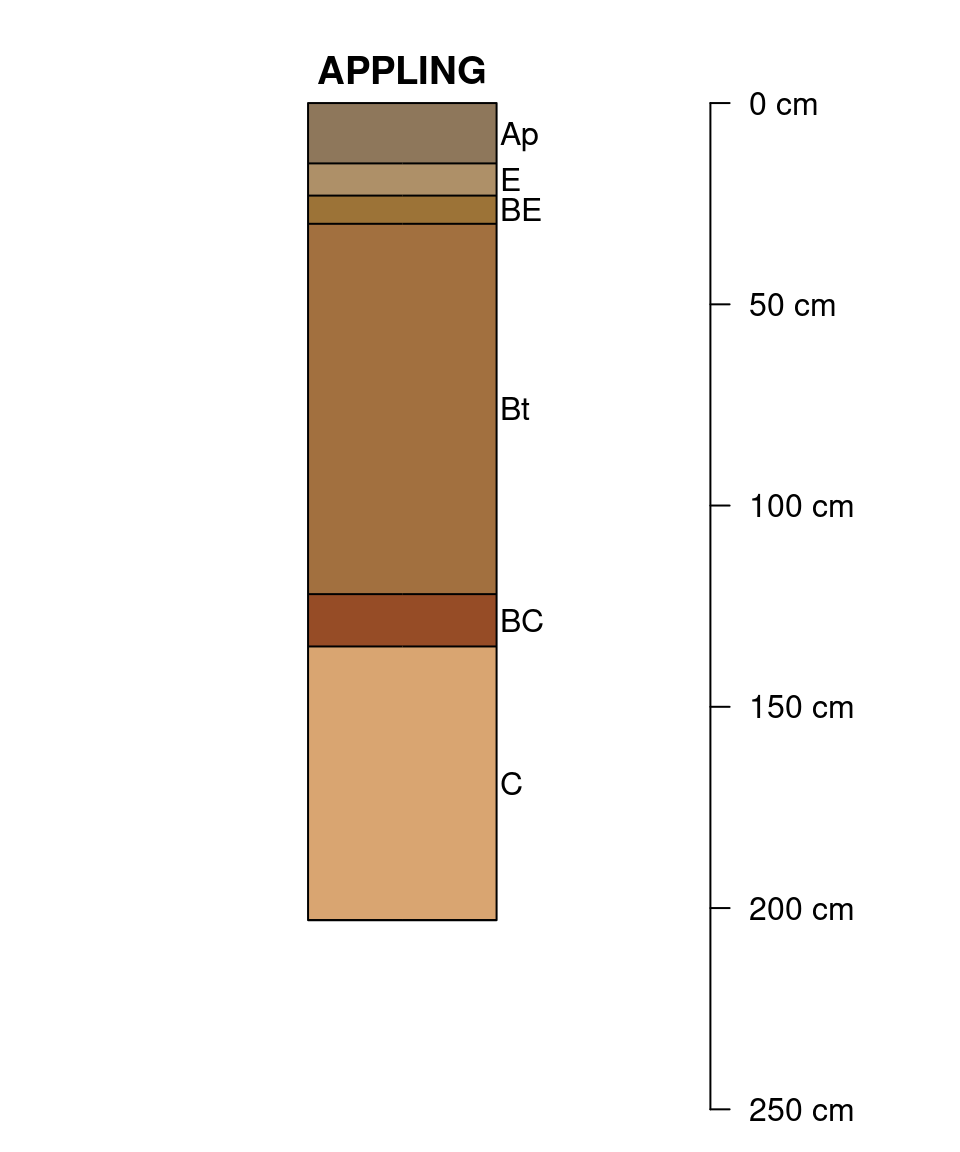
Small SoilProfileCollections
Making quality figures with fewer than 5 soil profiles usually
requires more customization of the basic call to plotSPC.
In general, the following are a good starting point:
- shrink margins and disable clipping with
par - adjust output graphic device (e.g.
png()) dimensions and resolution - increase font size with
cex.names - adjust sketch width with
width, typically within 0.15-0.35 - move depth axis to the left using negative
linevalues, e.g. (depth.axis = list(line = -2))
Get some example data from the Official Series Descriptions:
data(osd)
x <- osdUsing 5x6 inch output device:
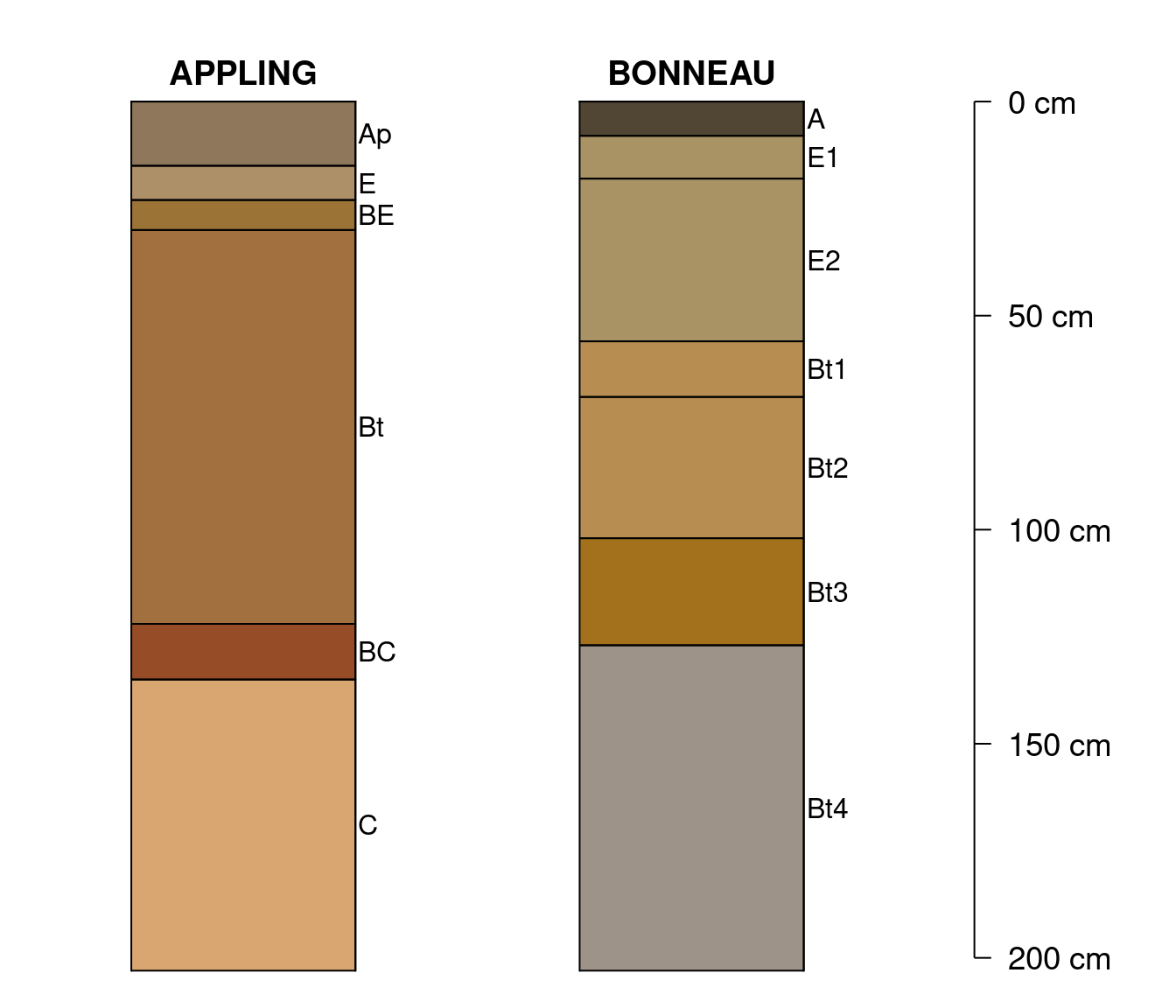
Using 7x6 inch output device, slight adjustments to
width usually required:
# set margins and turn off clipping
par(mar = c(0, 2, 0, 4), xpd = NA)
plotSPC(x[1:2, ], cex.names = 1, width = 0.25)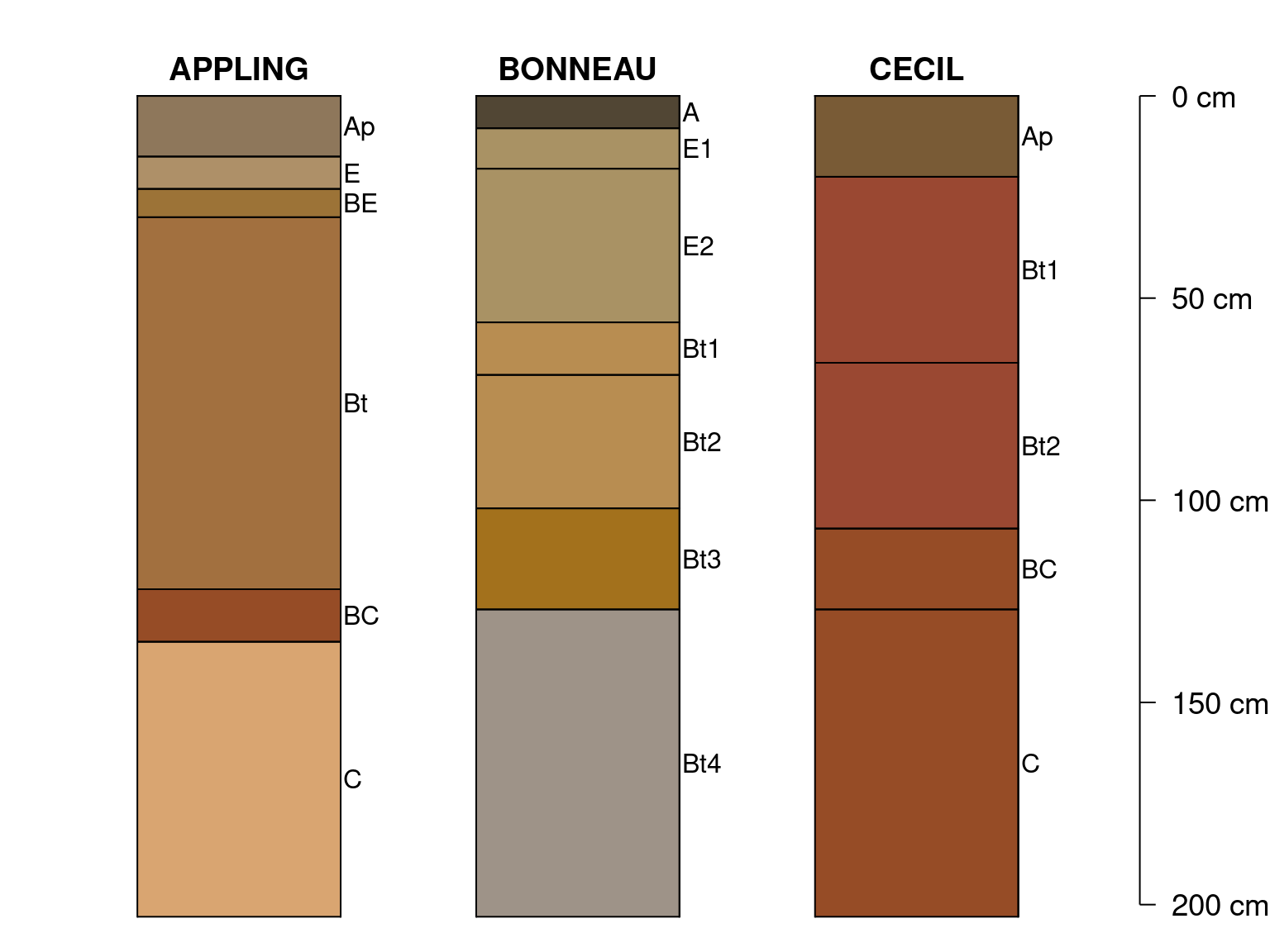
Using 8x6 inch output device, slight adjustments to are usually required:
par(mar = c(0, 2, 0, 4), xpd = NA)
plotSPC(x, cex.names = 1, depth.axis = list(line = -0.1), width = 0.3)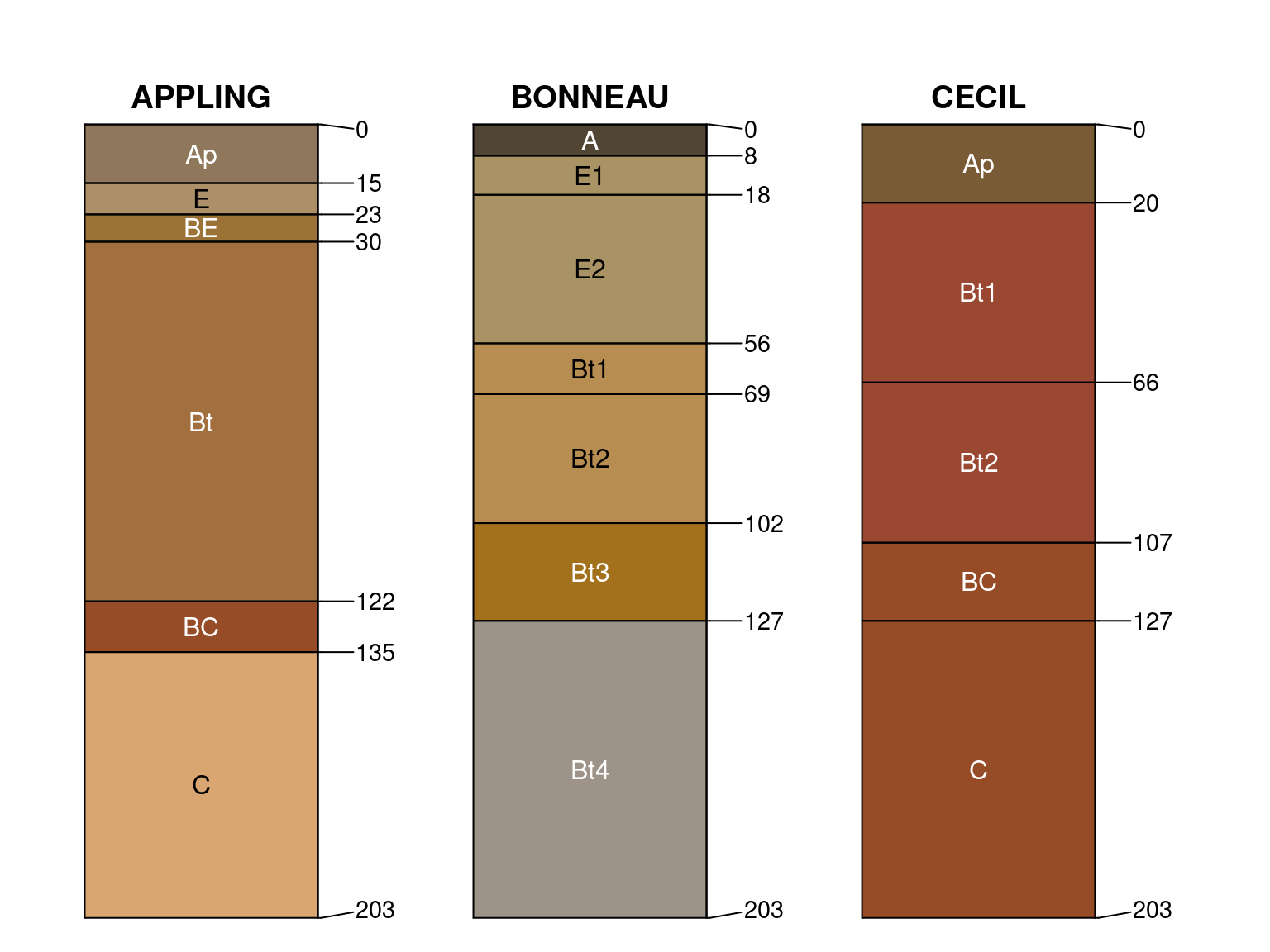
Horizon Depth Labeling
Horizon depths can be labeled for each profile as an alternative to a single depth axis.
par(mar = c(0, 0, 1, 1))
plotSPC(
x,
cex.names = 1,
name.style = 'center-center',
width = 0.3,
depth.axis = FALSE,
hz.depths = TRUE,
hz.depths.offset = 0.08
)
As of aqp version 1.41, it is possible to “fix” overlapping horizon
depth labels with the new fixLabelCollisions argument. This
approach to labeling depths works best when moving horizon designations
with the name.style argument. Note that setting
fixLabelCollisions = TRUE may incur a performance penalty
when plotting large SoilProfileCollection objects.
As of aqp version 2.0, fixLabelCollisions is set to
TRUE when setting hz.depths = TRUE.
Relative Horizontal Positioning
Relative horizontal positioning via relative.pos
argument. Can be used with plot.order but be careful:
relative.pos must be specified in the final
ordering of profiles. See ?plotSPC for details.
par(mar = c(4, 3, 2, 2))
pos <- jitter(1:length(sp4))
explainPlotSPC(sp4, name = 'name', relative.pos = pos)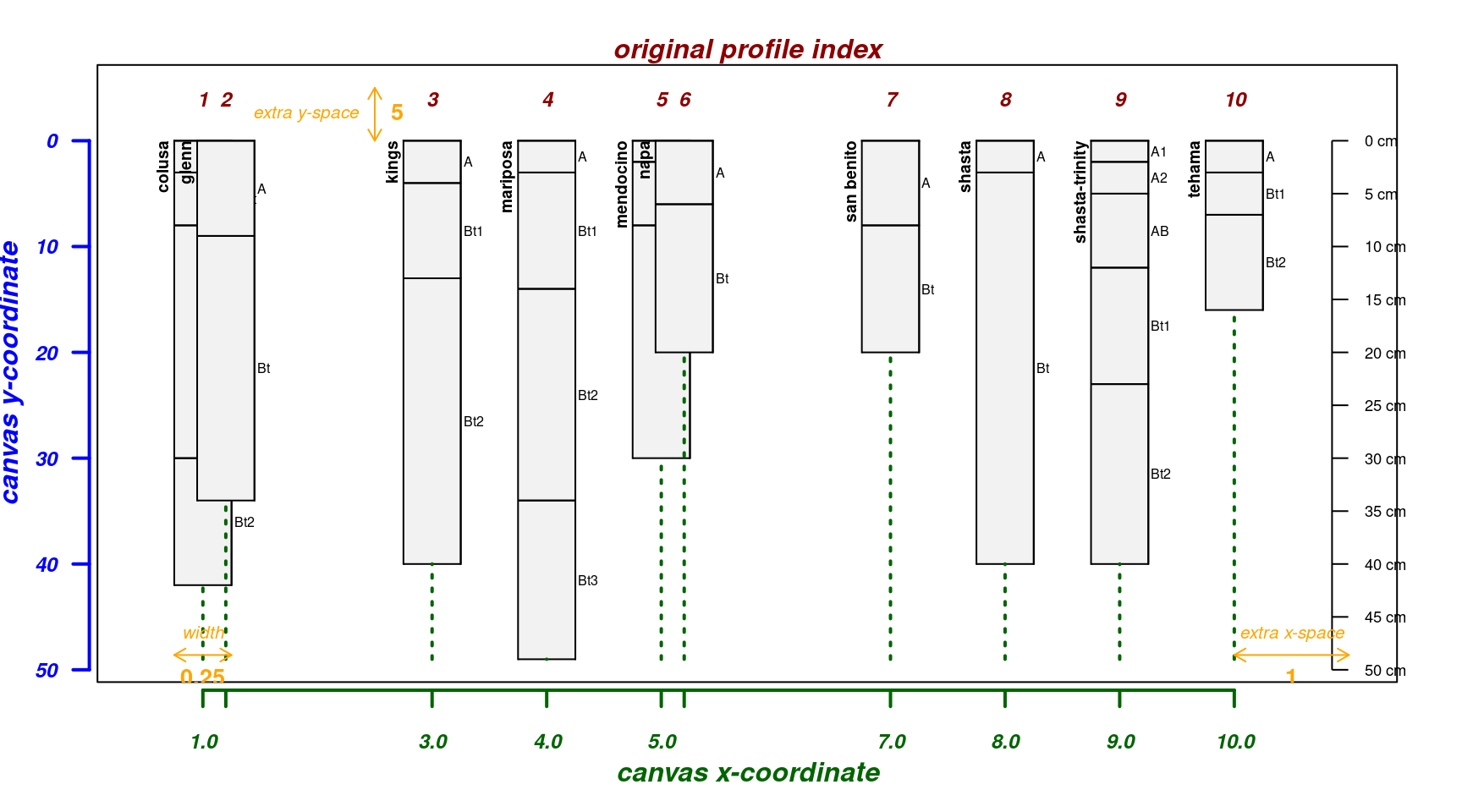
Relative positioning works well when the vector of positions is close to the default spacing along an integer sequence, but not when positions are closer than the width of a profile sketch.
par(mar = c(4, 3, 2, 2))
pos <- c(1, 1.2, 3, 4, 5, 5.2, 7, 8, 9, 10)
explainPlotSPC(sp4, name = 'name', relative.pos = pos)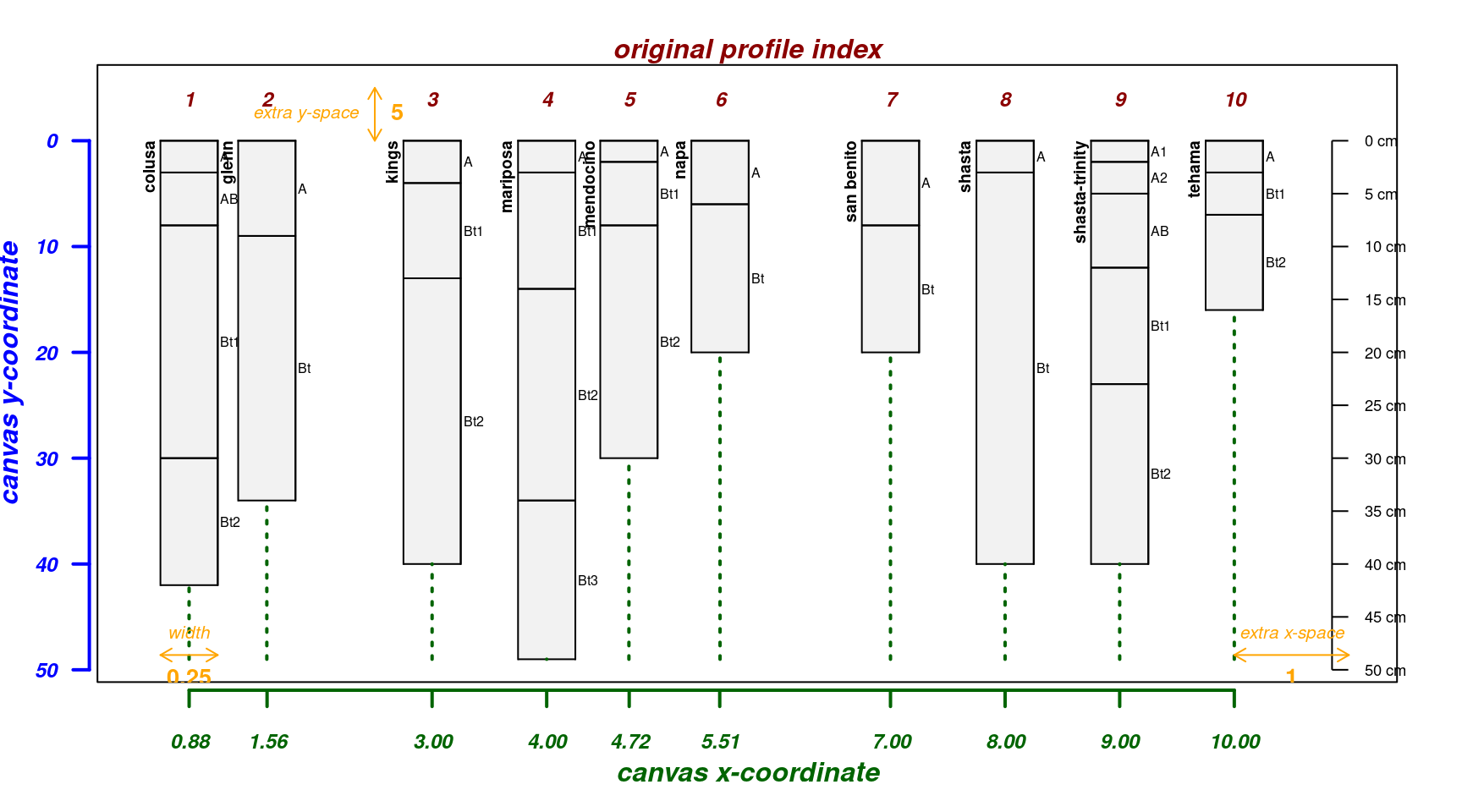
The fixOverlap() function can be used to find a suitable
arrangement of profiles based on a compromise between the suggested
relative positions and minimization of overlap.
This is an iterative procedure, based on random perturbations of
overlapping profiles, therefore it is possible for the algorithm to stop
at a sub-optimal configuration. Results can be controlled using
set.seed().
par(mar = c(4, 3, 2, 2))
new.pos <- fixOverlap(pos)
explainPlotSPC(sp4, name = 'name', relative.pos = new.pos)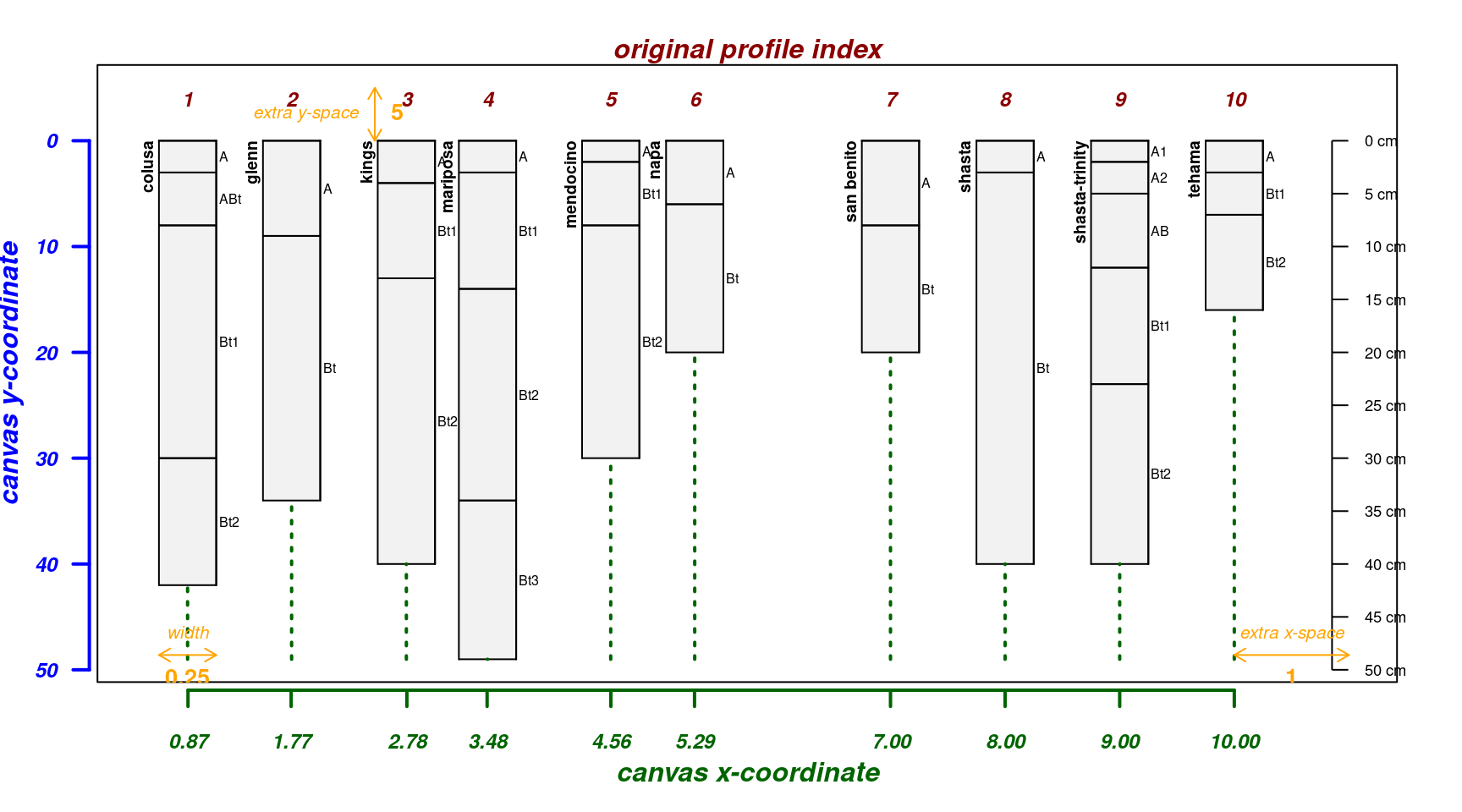
There are several parameters available for optimizing horizontal
position in the presence of overlap. See ?fixOverlap for
details and further examples.
par(mar = c(4, 3, 2, 2))
new.pos <- fixOverlap(pos, thresh = 0.7)
explainPlotSPC(sp4, name = 'name', relative.pos = new.pos)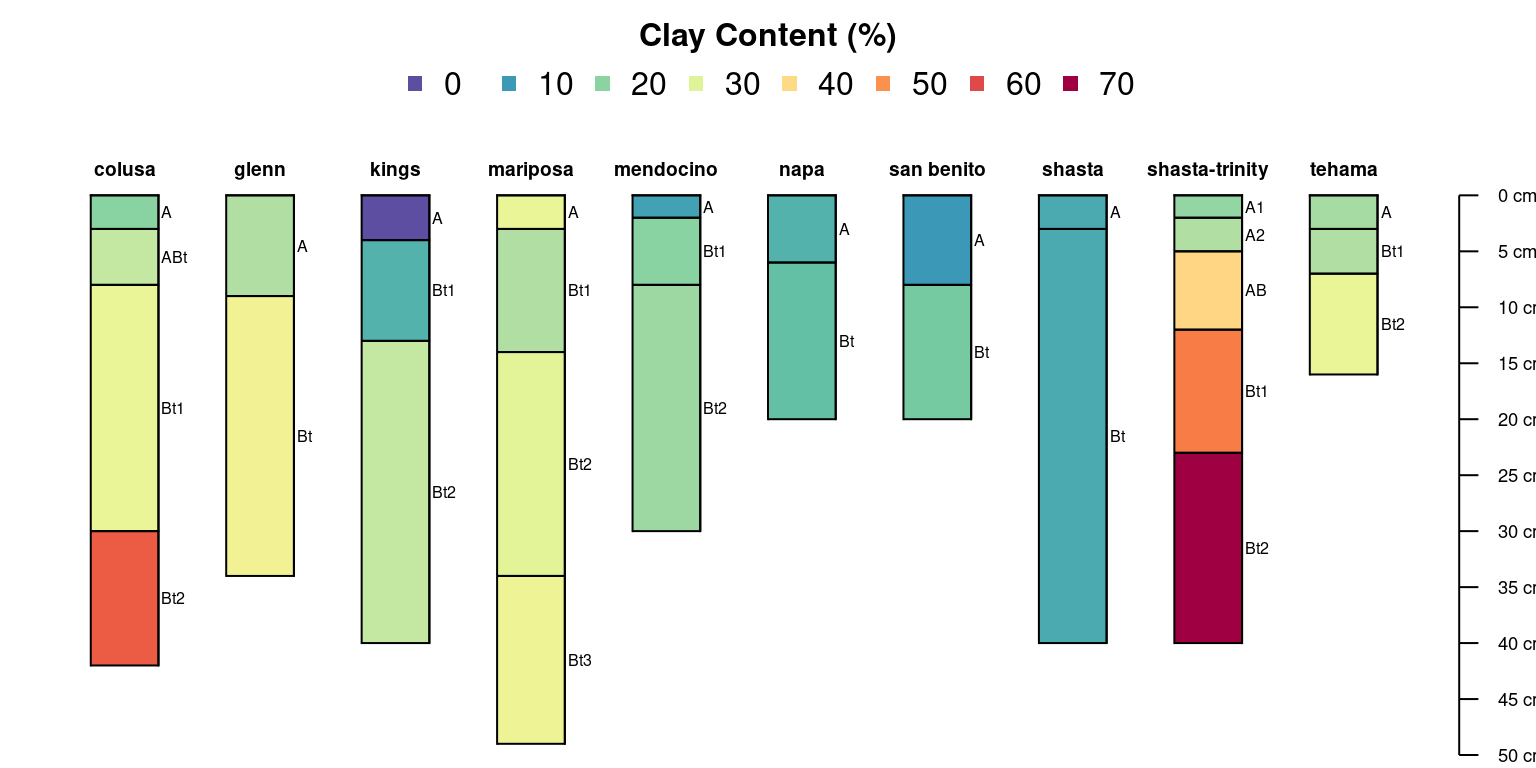 The SPC
plotting ideas tutorial contains several additional examples.
The SPC
plotting ideas tutorial contains several additional examples.
Thematic Sketches
Horizon-level attributes can be symbolized with color, in this case using the horizon-level attribute “clay”:
par(mar = c(0, 0, 3, 0))
plotSPC(sp4,
name = 'name',
color = 'clay',
col.label = 'Clay Content (%)')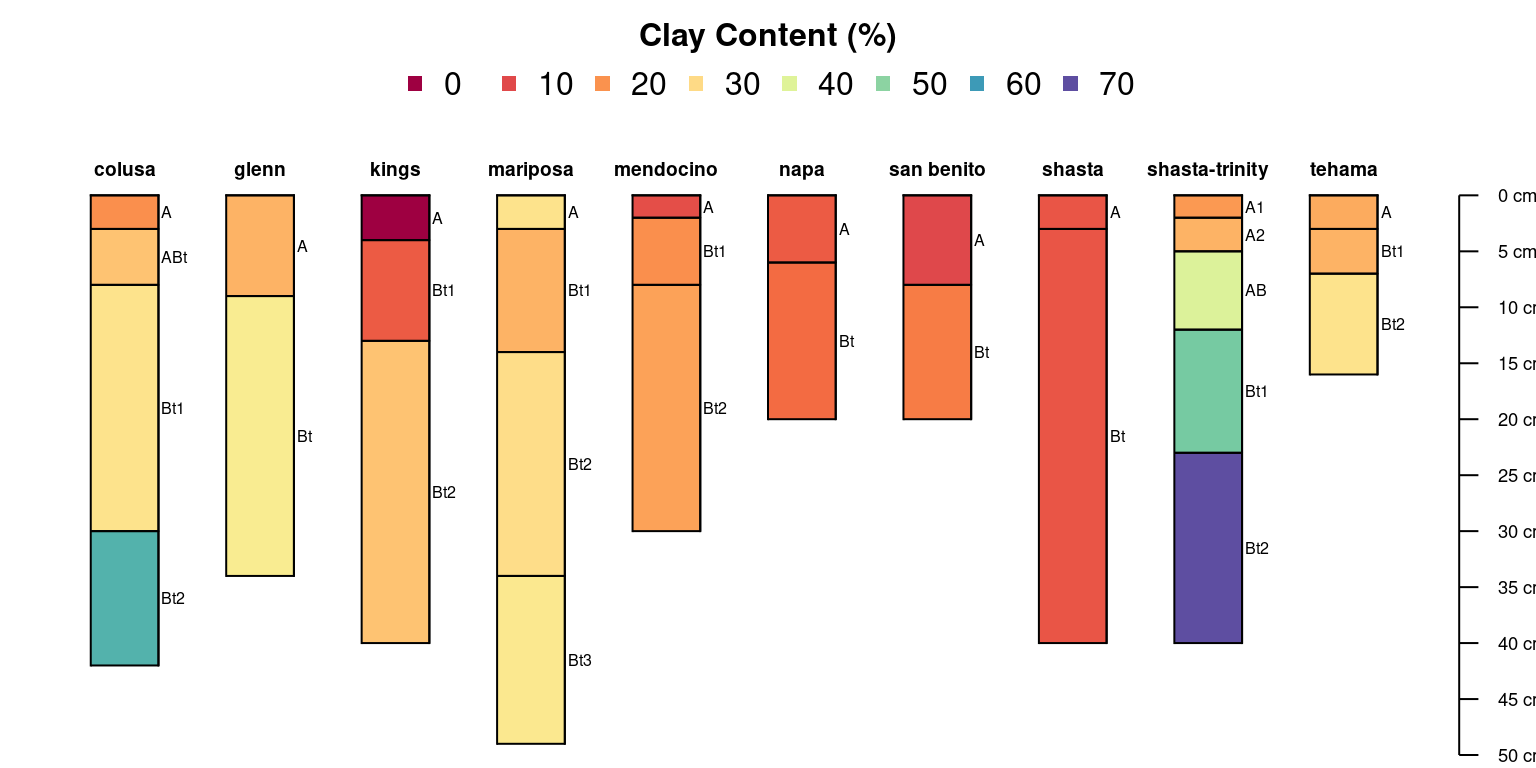
Use a different set of colors:
par(mar = c(0, 0, 3, 0))
plotSPC(
sp4,
name = 'name',
color = 'clay',
col.label = 'Clay Content (%)'
)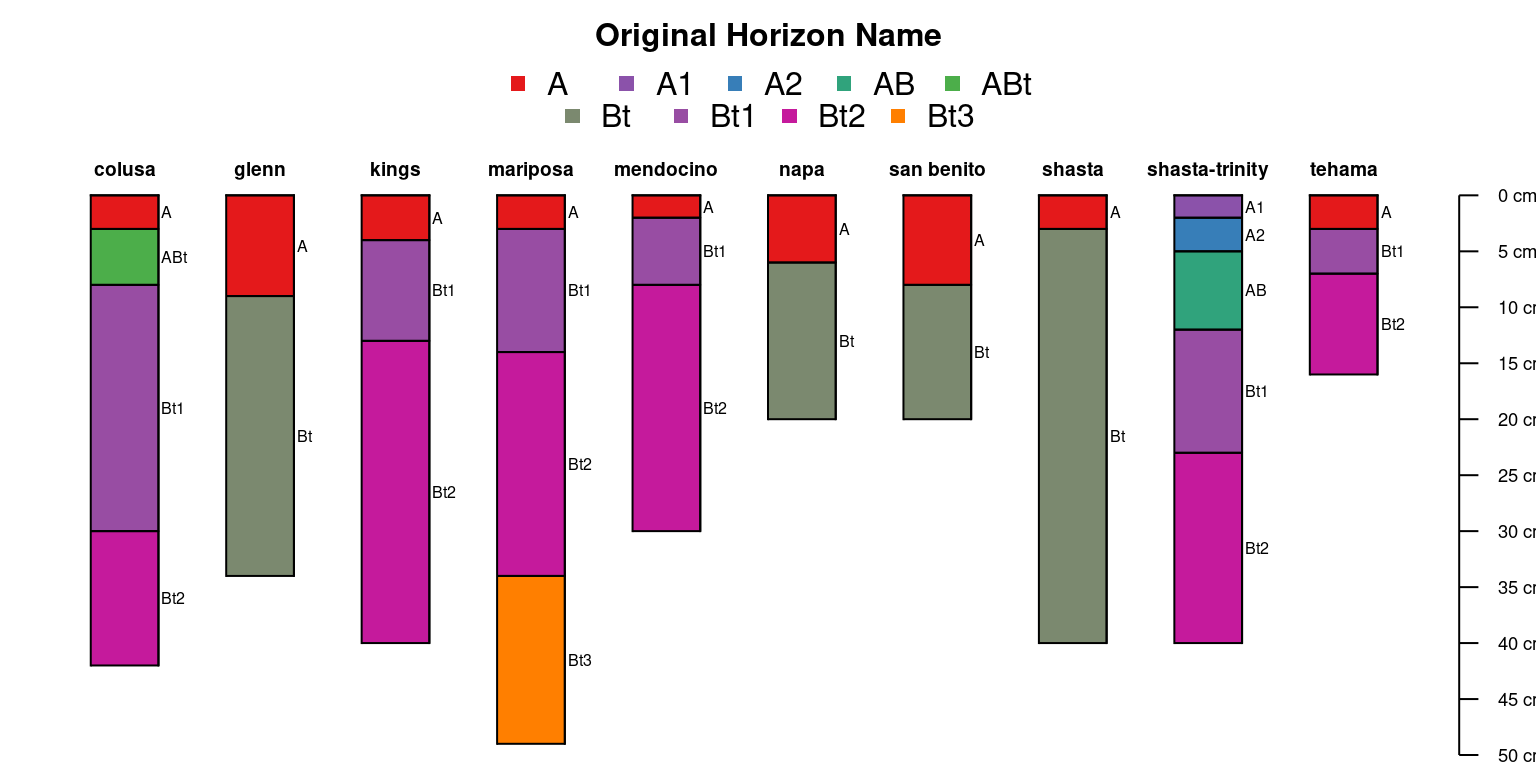
Categorical properties can also be used to make a thematic sketch.
Colors are interpolated when there are more classes than colors provided
by col.palette:
par(mar = c(0, 0, 3, 0))
plotSPC(
sp4,
name = 'name',
color = 'name',
col.label = 'Original Horizon Name'
)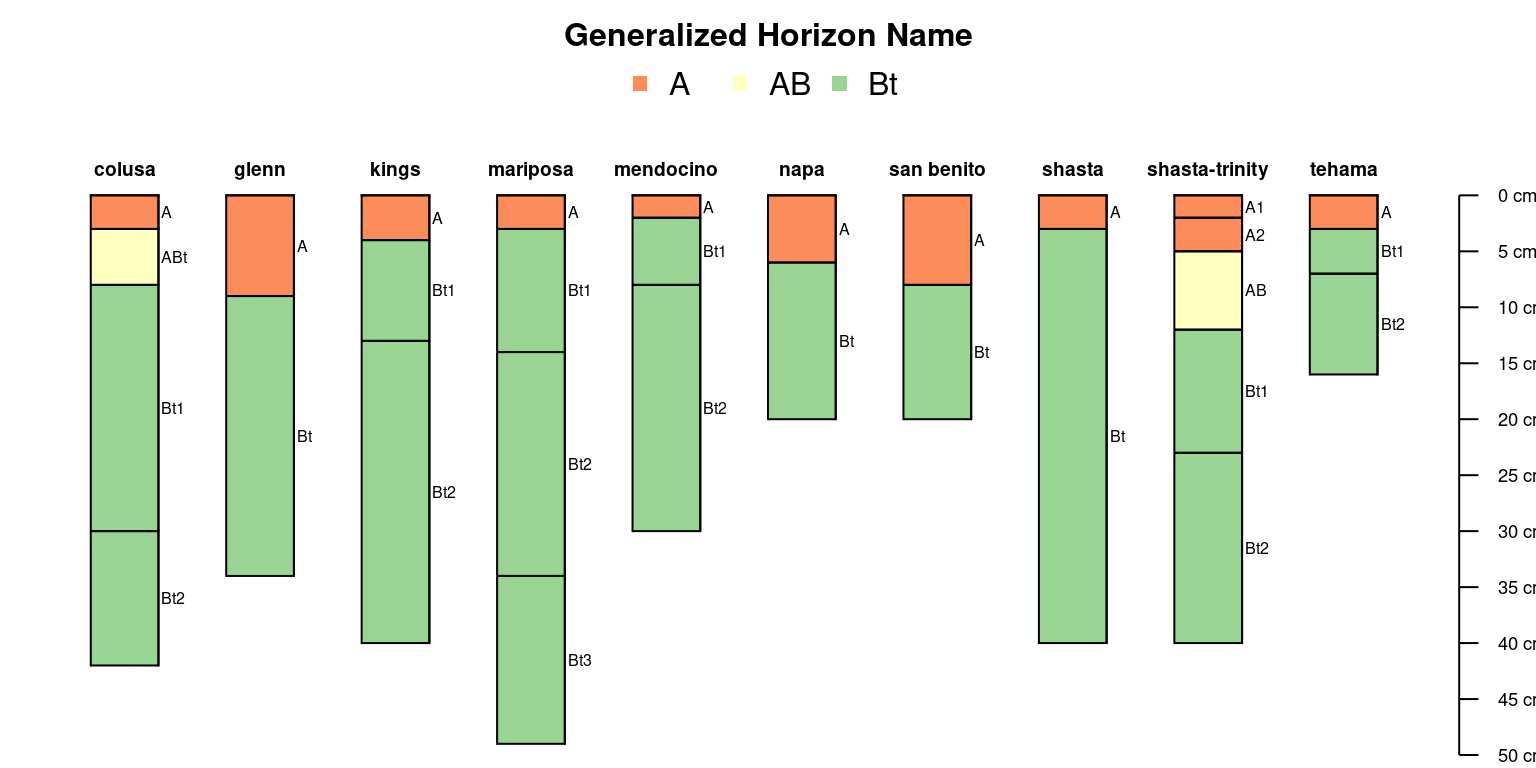
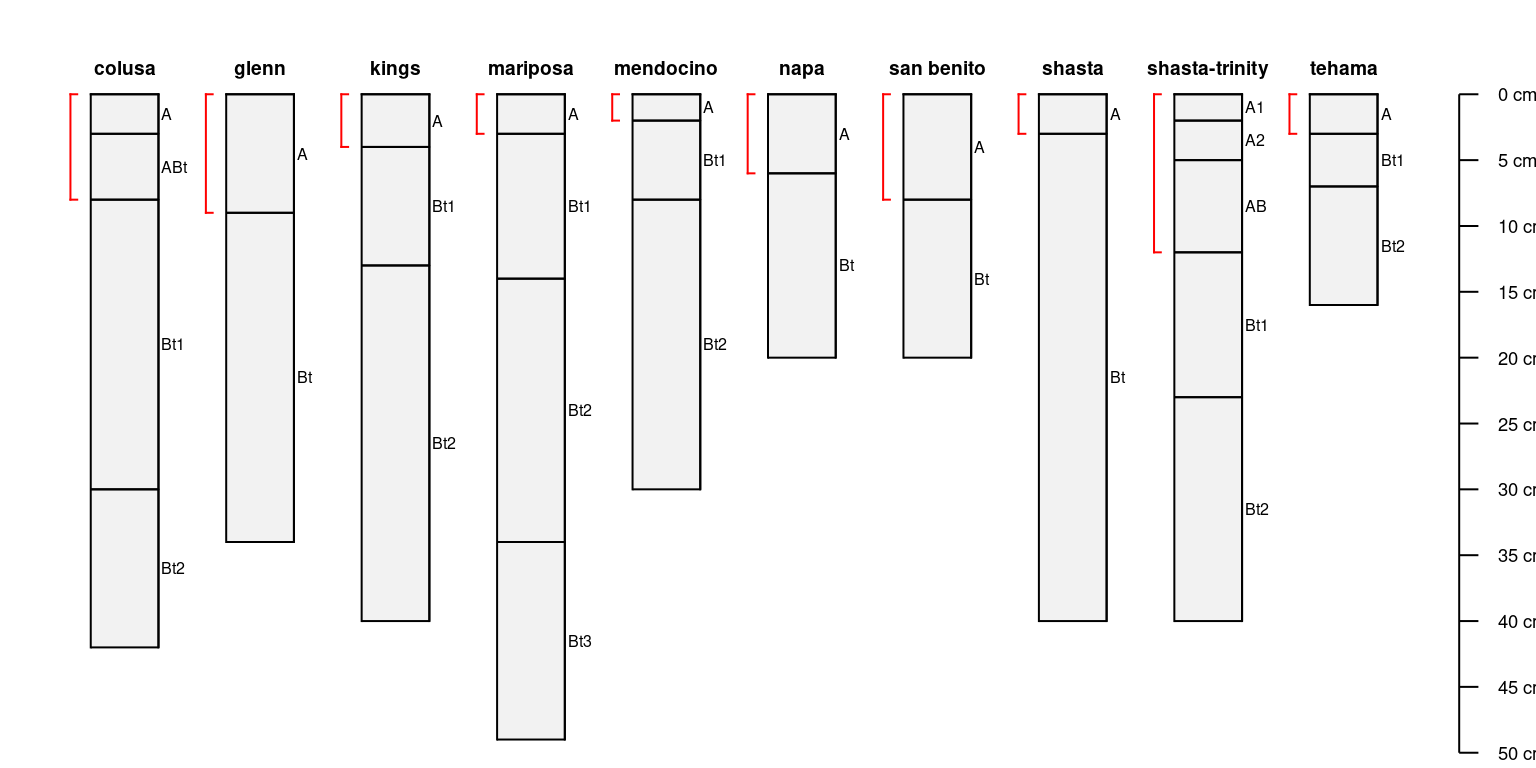
Try with generalized horizon labels:
par(mar = c(0, 0, 3, 0))
# generalize horizon names into 3 groups
sp4$genhz <- generalize.hz(sp4$name, new = c('A', 'AB', 'Bt'), pat = c('A[0-9]?', 'AB', '^Bt'))
plotSPC(
sp4,
name = 'name',
color = 'genhz',
col.label = 'Generalized Horizon Name'
)
Horizon-level attributes that represent a volume fraction
(e.g. coarse-fragment percentage) can be added to an existing figure.
See ?addVolumeFraction for adding layers to an existing
plot based on these attributes.
par(mar = c(0, 0, 3, 0))
# convert coarse rock fragment proportion to percentage
sp4$frag_pct <- sp4$CF * 100
# label horizons with fragment percent
plotSPC(sp4, name = 'frag_pct', color = 'frag_pct')
# symbolize volume fraction data
addVolumeFraction(sp4, colname = 'frag_pct')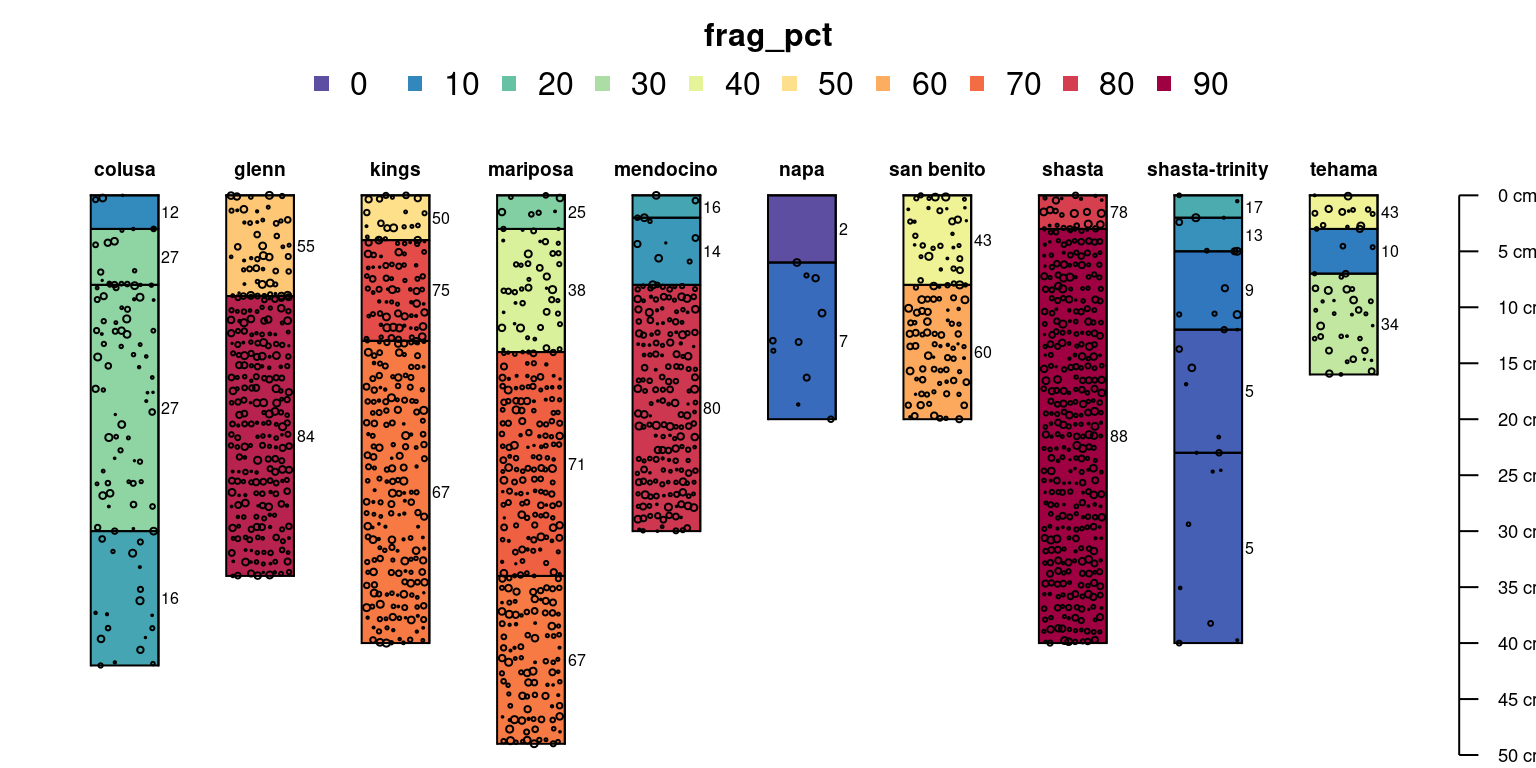
Depth Intervals
Annotation of depth-intervals can be accomplished using “brackets”.
Here we extract top/bottom depths associated with top and bottom depth A
horizons using the min/max variants of the depthOf()
function. See ?depthOf for more details.
# extract top/bottom depths associated with all A horizons
tops <- minDepthOf(sp4, pattern = '^A', hzdesgn = 'name', top = TRUE)
bottoms <- maxDepthOf(sp4, pattern = '^A', hzdesgn = 'name', top = FALSE)
IDs <- profile_id(sp4)
# assemble data.frame
a <- data.frame(id = IDs, top = tops$top, bottom = bottoms$bottom)
# check
a#> id top bottom
#> 1 colusa 0 8
#> 2 glenn 0 9
#> 3 kings 0 4
#> 4 mariposa 0 3
#> 5 mendocino 0 2
#> 6 napa 0 6
#> 7 san benito 0 8
#> 8 shasta 0 3
#> 9 shasta-trinity 0 12
#> 10 tehama 0 3
par(mar = c(0, 0, 0, 0))
plotSPC(sp4)
# annotate A horizon depth interval with brackets
addBracket(a, col = 'red', offset = -0.4)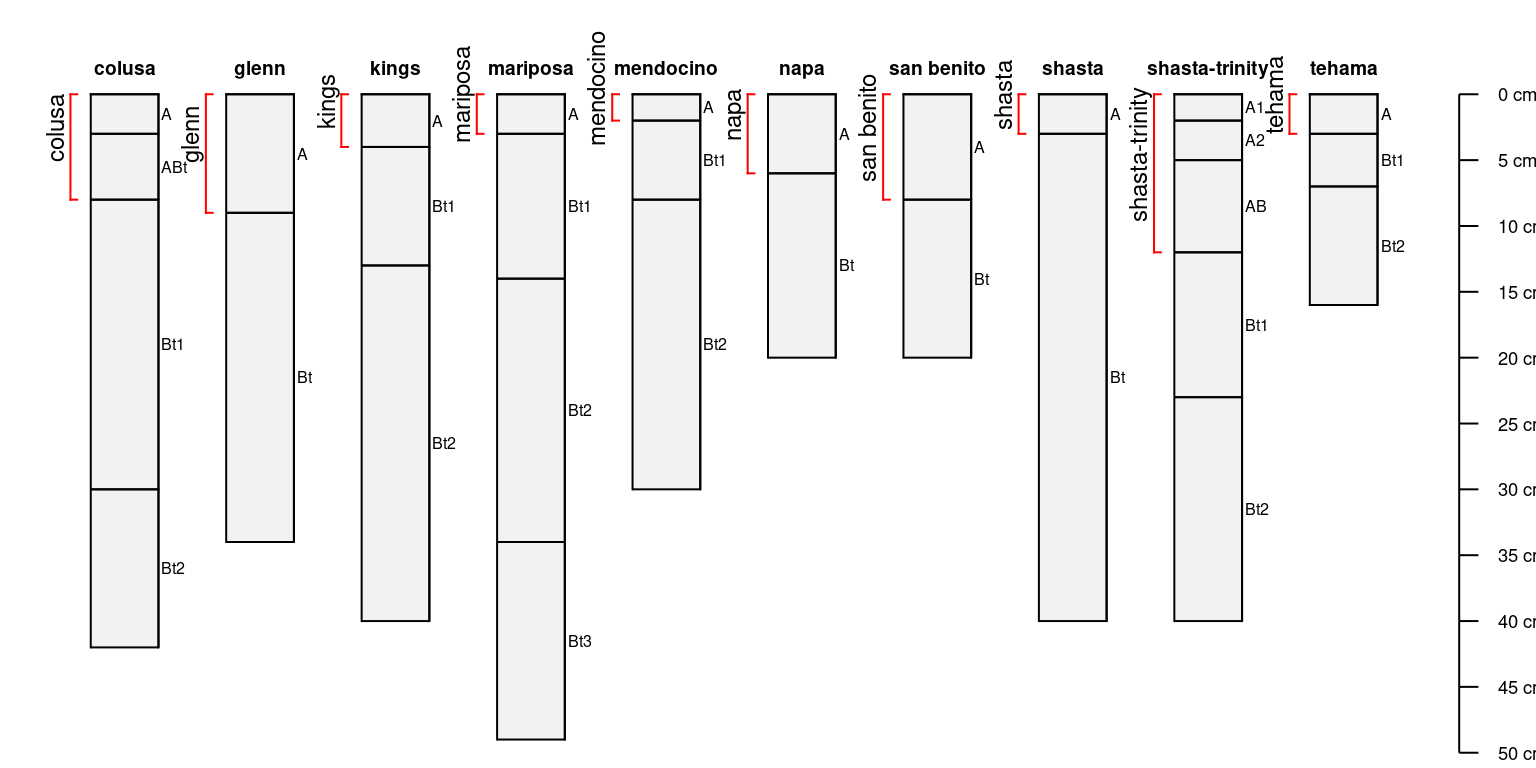
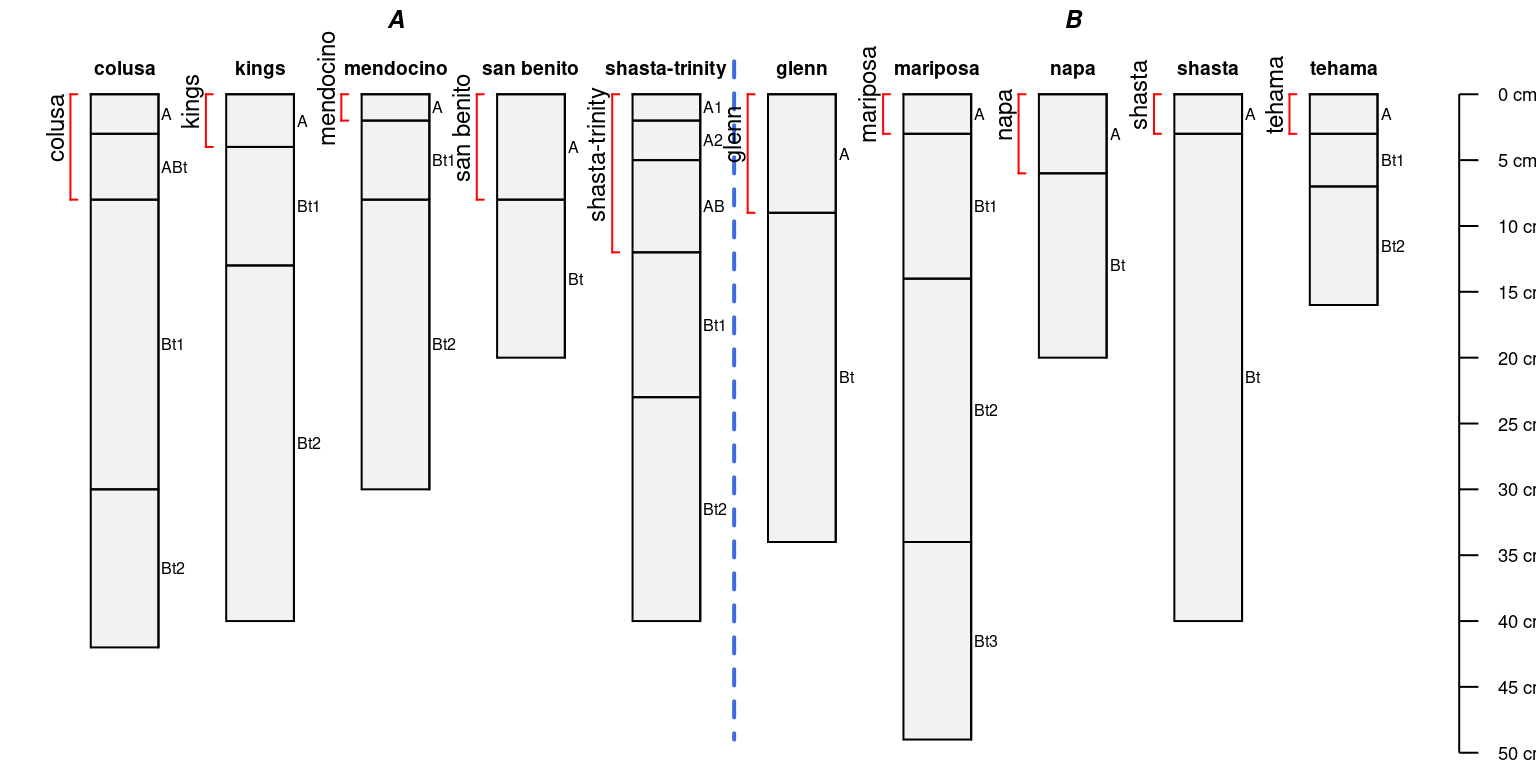
Add labels:
par(mar = c(0, 0, 0, 0))
plotSPC(sp4, name = 'name')
# label
a$label <- site(sp4)$id
# note that depth brackets "know which profiles to use" via profile ID
addBracket(
a,
labcol = 'label',
col = 'red',
label.cex = 0.75,
missing.bottom.depth = 25,
offset = -0.4
)
It is possible to arrange profile sketches by site-level grouping variable:
par(mar = c(0, 0, 0, 0))
groupedProfilePlot(sp4, groups = 'group')
addBracket(a, col = 'red', offset = -0.4)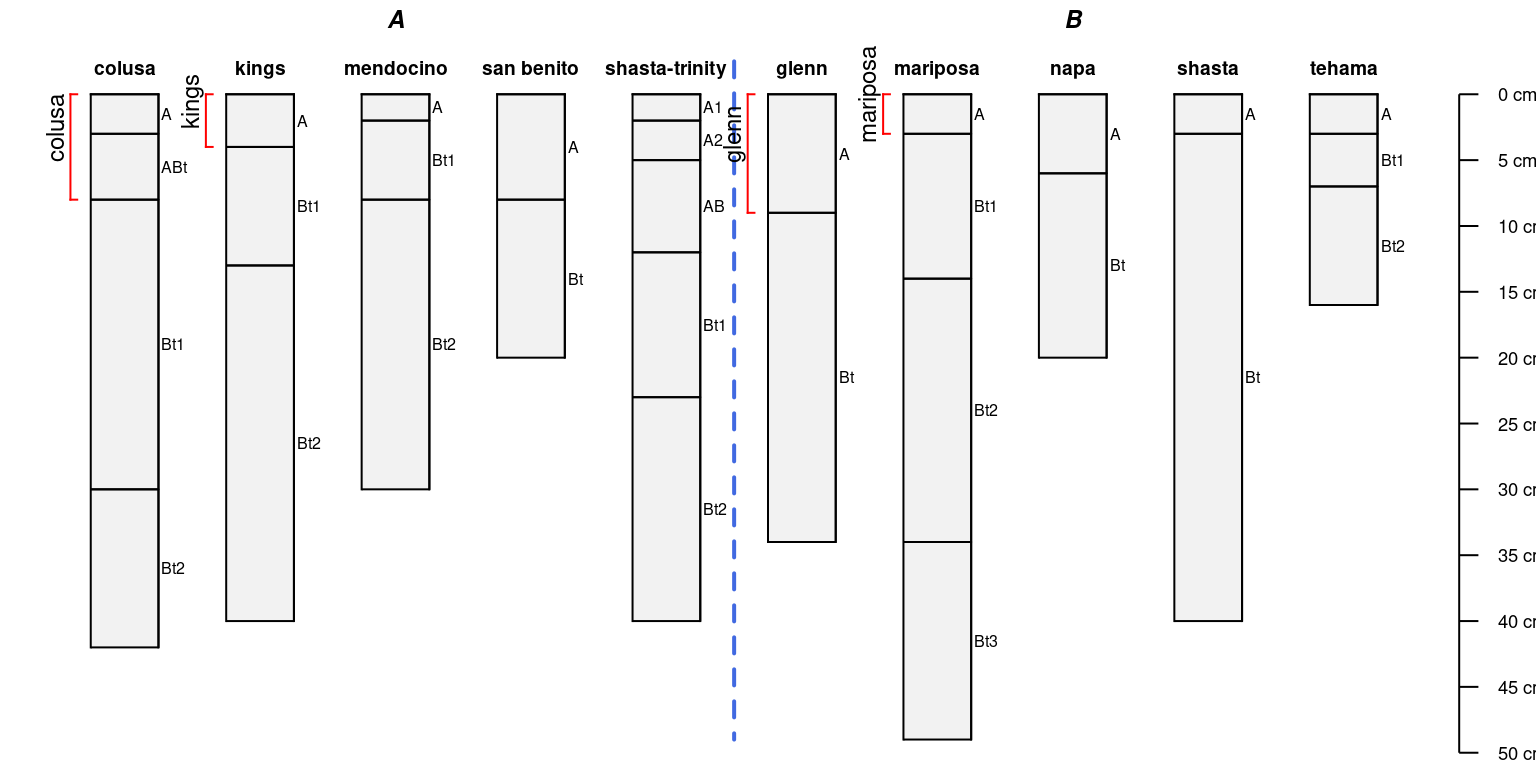
There need not be brackets for all profiles in a collection:
par(mar = c(0, 0, 0, 0))
a.sub <- a[1:4,]
groupedProfilePlot(sp4, groups = 'group')
addBracket(a.sub, col = 'red', offset = -0.4)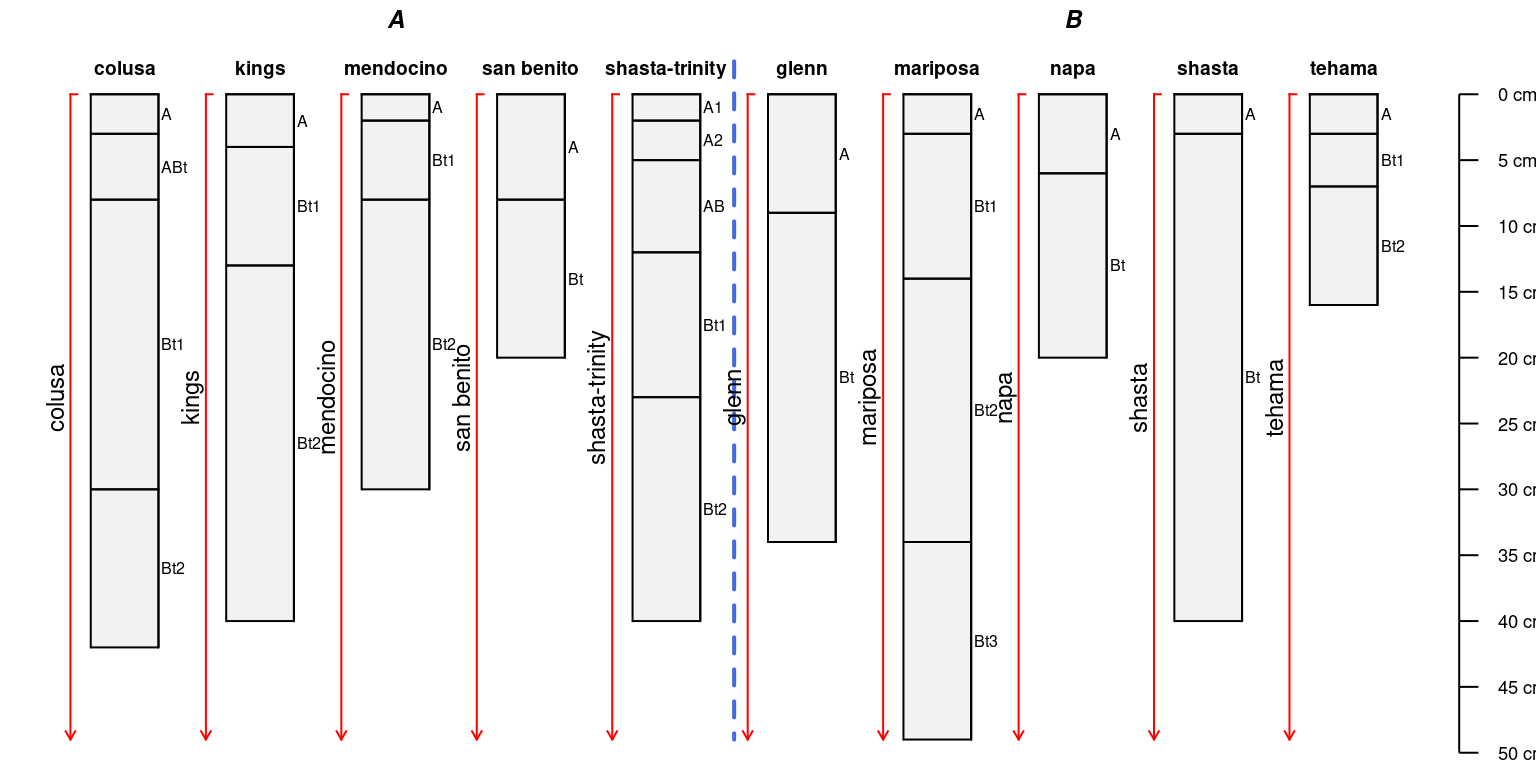
When bottom depths are missing an arrow is used:
a$bottom <- NA
par(mar = c(0, 0, 0, 0))
groupedProfilePlot(sp4, groups = 'group')
addBracket(a, col = 'red', offset = -0.4)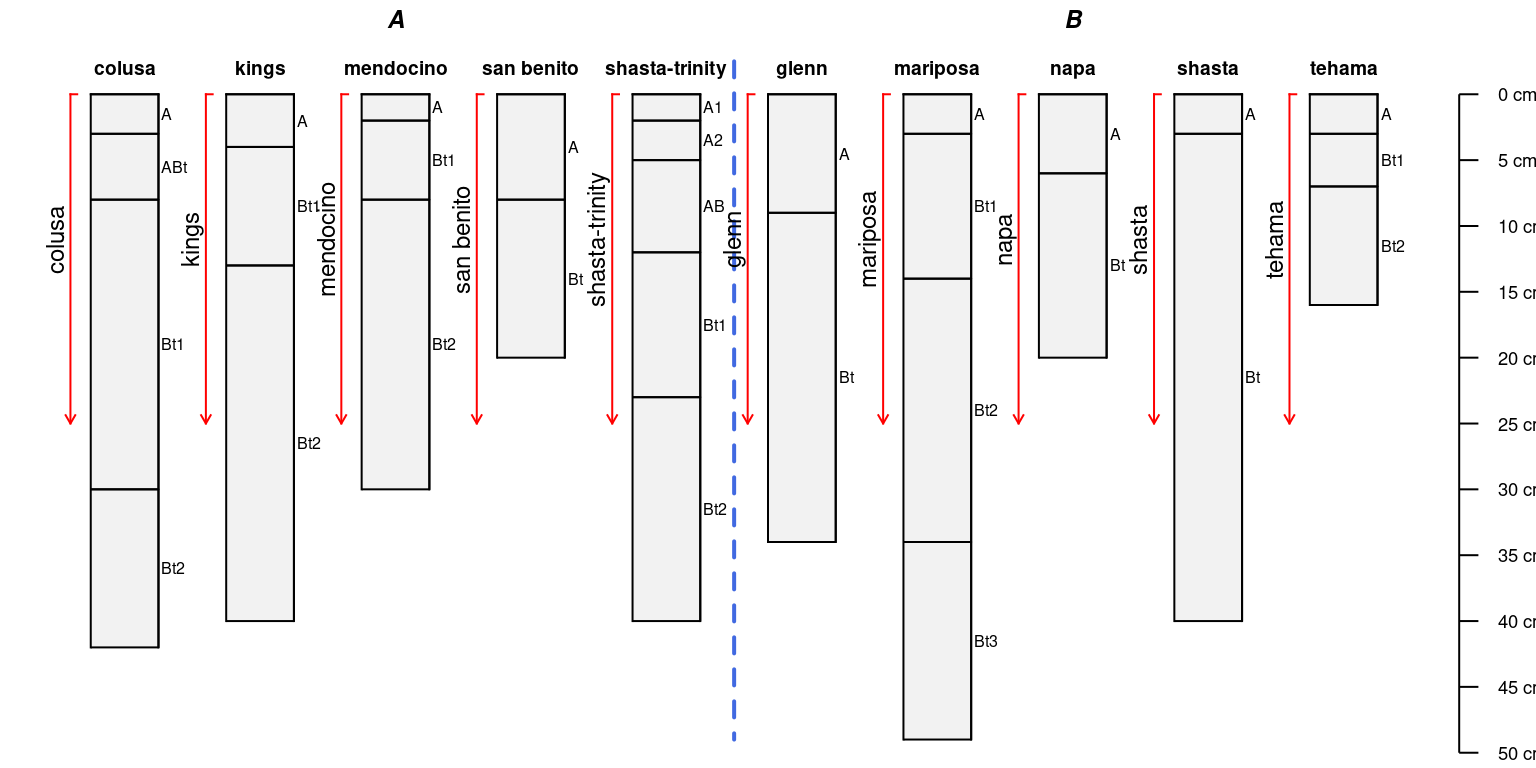
Manually define bottom depth:
par(mar = c(0, 0, 0, 0))
groupedProfilePlot(sp4, groups = 'group')
addBracket(
a,
col = 'red',
label.cex = 0.75,
missing.bottom.depth = 25,
offset = -0.4
)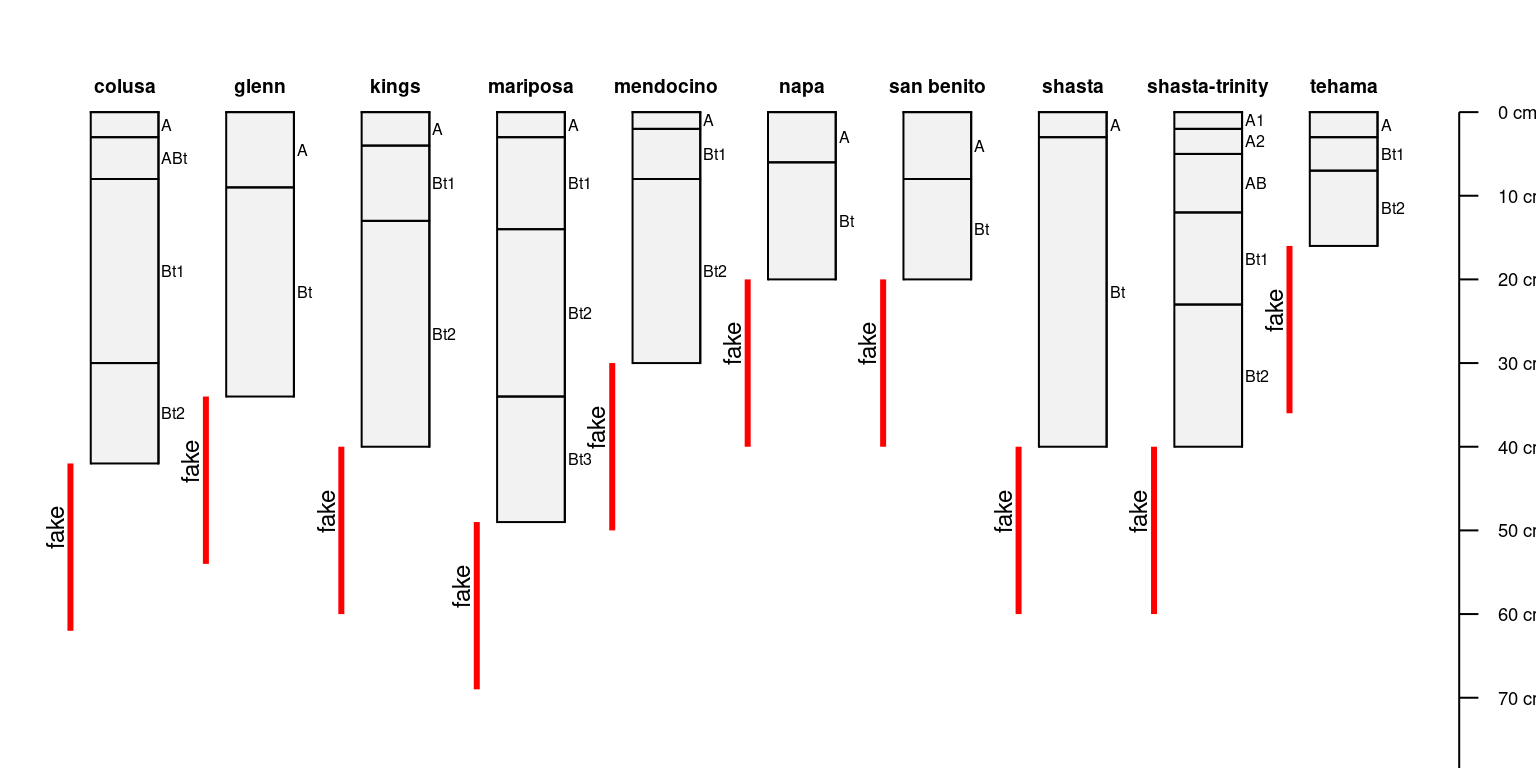
Further customization of brackets:
par(mar = c(0, 0, 0, 0))
plotSPC(sp4, max.depth = 75)
# copy root-restricting features
a <- restrictions(sp4)
# add a label: restrictive feature 'kind'
a$label <- a$kind
# add restrictions using vertical bars
addBracket(
a,
labcol = 'label',
col = 'red',
label.cex = 0.75,
tick.length = 0,
lwd = 3,
offset = -0.4
)
Notes
These functions (addBracket(),
addDiagnosticBracket(), and
addVolumeFraction()) will automatically compensate for
alternative sketch ordering or relative positioning of profiles.
SVG Output for Use in Page Layout Tools
Sketches for use in layout tools such as Adobe Illustrator or
Inkscape should be stored in a vector file format. SVG is compatible
with most software titles. WMF output is compatible with MS Office tools
and can be written with the win.metafile() function.
# library(svglite)
# svglite(filename = 'e:/temp/fig.svg', width = 7, height = 6, pointsize = 12)
#
# par(mar = c(0,2,0,4), xpd = NA)
# plotSPC(x, cex.names=1, depth.axis = list(line = -0.2), width=0.3)
#
# dev.off()Iterating Over Profiles in a Collection
The profileApply() function is an extension of the
familiar apply() family of functions extended to
SoilProfileCollection objects.
The function named in the FUN argument is evaluated once
for each profile in the collection, typically returning a single value
per profile. In this case, the ordering of the results would match the
ordering of values in the site level attribute table.
# max() returns the depth of a soil profile
sp4$soil.depth <- profileApply(sp4, FUN = max)
# max() with additional argument give max depth to non-missing 'clay'
sp4$soil.depth.clay <- profileApply(sp4, FUN = max, v = 'clay')
# nrow() returns the number of horizons
sp4$n.hz <- profileApply(sp4, FUN = nrow)
# compute the mean clay content by profile using an inline function
sp4$mean.clay <- profileApply(sp4, FUN = function(i) mean(i$clay))
# estimate soil depth based on horizon designation
sp4$soil.depth <- profileApply(sp4, estimateSoilDepth, name = 'name')When FUN returns a vector of the same length as the
number of horizons in a profile, profileApply() can be used
to create new horizon-level attributes. For example, the change in clay
content by horizon depth (delta.clay, below) could be
calculated as:
# save as horizon-level attribute
sp4$delta.clay <- profileApply(sp4, FUN = function(i) c(NA, diff(i$clay)))
# check results:
horizons(sp4)[1:6, c('id', 'top', 'bottom', 'clay', 'delta.clay')]#> id top bottom clay delta.clay
#> 1 colusa 0 3 21 NA
#> 2 colusa 3 8 27 6
#> 3 colusa 8 30 32 5
#> 4 colusa 30 42 55 23
#> 5 glenn 0 9 25 NA
#> 6 glenn 9 34 34 9More complex summaries can be generated by writing a custom function
that is then called by profileApply(). Note that each
profile is passed into this function and accessed via a temporary
variable (i), which is a SoilProfileCollection
object containing a single profile. See help(profileApply)
for details.
A list of SoilProfileCollection objects returned from a
custom function can be combined into a single
SoilProfileCollection object via
combine().
# compute hz-thickness weighted mean exchangeable-Ca:Mg
wt.mean.ca.mg <- function(i) {
# use horizon thickness as a weight
thick <- i$bottom - i$top
# compute the thickness weighted mean, ignoring missing values
weighted.mean(i$ex_Ca_to_Mg, w = thick, na.rm = TRUE)
}
# apply our custom function and save results as a site-level attribute
sp4$wt.mean.ca.to.mg <- profileApply(sp4, wt.mean.ca.mg)We can now use our some of our new site-level attributes to order the profiles when plotting.
In this case profiles are ordered based on the horizon-thickness weighted mean, exchangeable Ca:Mg values. Horizons are colored by exchangeable Ca:Mg values.
# plot the data using our new order based on Ca:Mg weighted average
# the result is an index of rank
new.order <- order(sp4$wt.mean.ca.to.mg)
par(mar = c(4, 0, 3, 0)) # tighten figure margins
plotSPC(sp4,
name = 'name',
color = 'ex_Ca_to_Mg',
plot.order = new.order)
# add an axis labeled with the sorting criteria
axis(1, at = 1:length(sp4), labels = round(sp4$wt.mean.ca.to.mg, 3), cex.axis = 0.75)
mtext(1, line = 2.25, text = 'Horizon Thickness Weighted Mean Ex. Ca:Mg', cex = 0.75)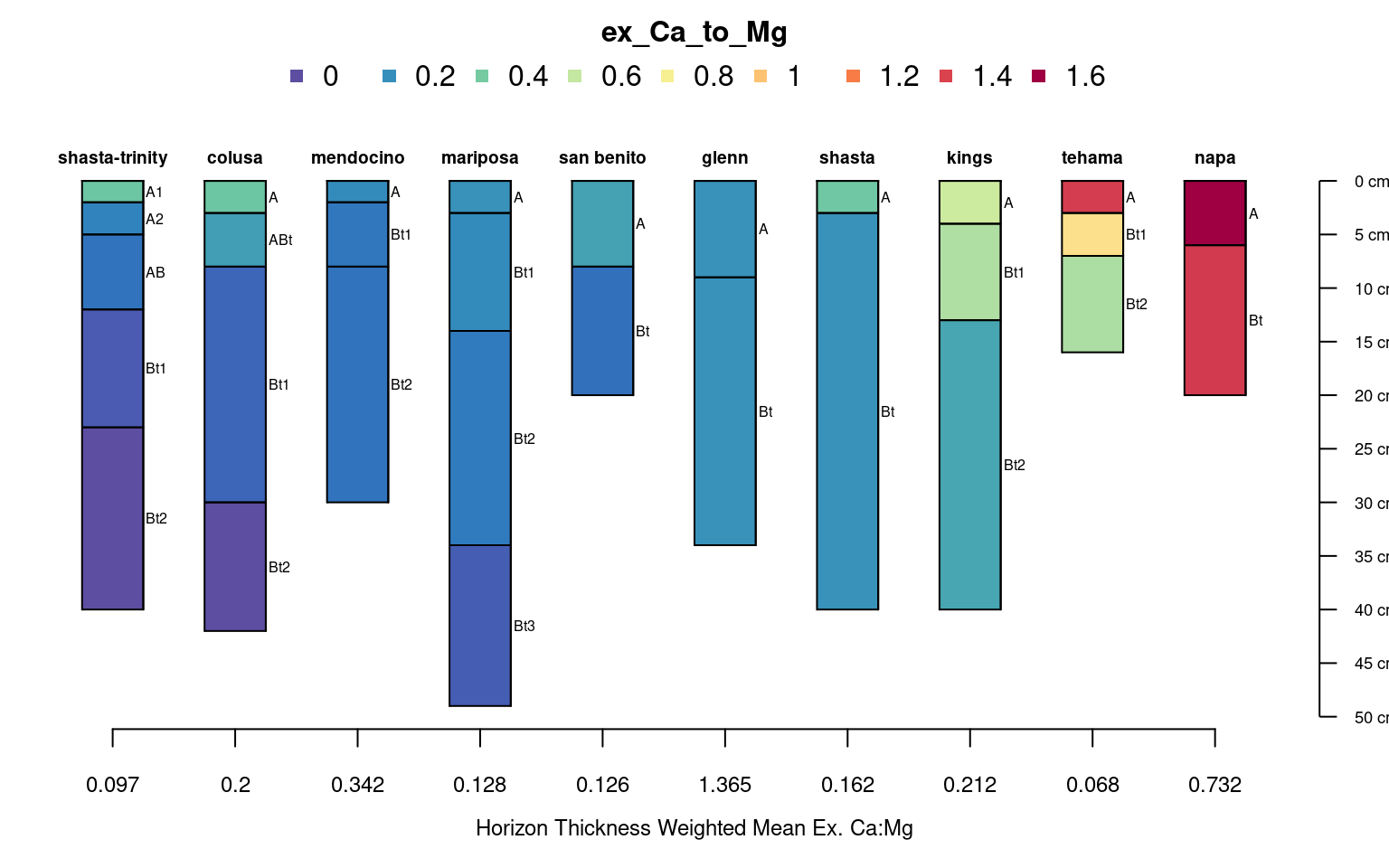
Depth-wise Selection and Resampling
Slicing Horizons: dice()
Collections of soil profiles can be sliced (or re-sampled) into
regular depth-intervals with the dice() function. The
slicing structure and variables of interest are defined via formula
notation:
# slice select horizon-level attributes
seq ~ var.1 + var.2 + var.3 + ...
# slice all horizon-level attributes
seq ~ .where seq is a sequence of integers
(e.g. 0:15, c(5,10,15,20), etc.) and
var.1 + var.2 + var.3 + ... are horizon-level attributes to
slice. Both continuous and categorical variables can be named on the
right-hand-side of the formula. The results returned by
dice() is either a SoilProfileCollection, or
data.frame when called with the optional argument
SPC = FALSE. For example, to slice our sample data set into
1-cm intervals, from 0–15 cm:
# resample to 1cm slices
s <- dice(sp4, fm = 0:15 ~ sand + silt + clay + name + ex_Ca_to_Mg)
# check the result
class(s)#> [1] "SoilProfileCollection"
#> attr(,"package")
#> [1] "aqp"
# plot sliced data
par(mar = c(0, 0, 3, 0)) # tighten figure margins
plotSPC(s, color = 'ex_Ca_to_Mg')
Once soil profile data have been sliced, it is simple to extract
“chunks” of data by depth interval via [-subsetting:
# slice from 0 to max depth in the collection
s <- dice(sp4, fm= 0:max(sp4) ~ sand + silt + clay + name + ex_Ca_to_Mg)
# extract all data over the range of 5--10 cm:
plotSPC(s[, 5:10])
# extract all data over the range of 25--50 cm:
plotSPC(s[, 25:50])
# extract all data over the range of 10--20 and 40--50 cm:
plotSPC(s[, c(10:20, 40:50)])Ragged Extraction: glom()
Select all horizons that overlap the interval of 5-15cm:
# truncate to the interval 5-15cm
clods <- glom(sp4, z1 = 5, z2 = 15)
# plot outlines of original profiles
par(mar = c(0, 0, 3, 1.5))
plotSPC(sp4, color = NA, name = NA, print.id = FALSE, depth.axis = FALSE, lwd = 0.5)
# overlay glom() depth criteria
rect(xleft = 0.5, ybottom = 15, xright = length(sp4) + 0.5, ytop = 5, lty = 2)
# add SoilProfileCollection with selected horizons
plotSPC(clods, add = TRUE, cex.names = 0.6, name = 'name', color = 'ex_Ca_to_Mg', name.style = 'center-center')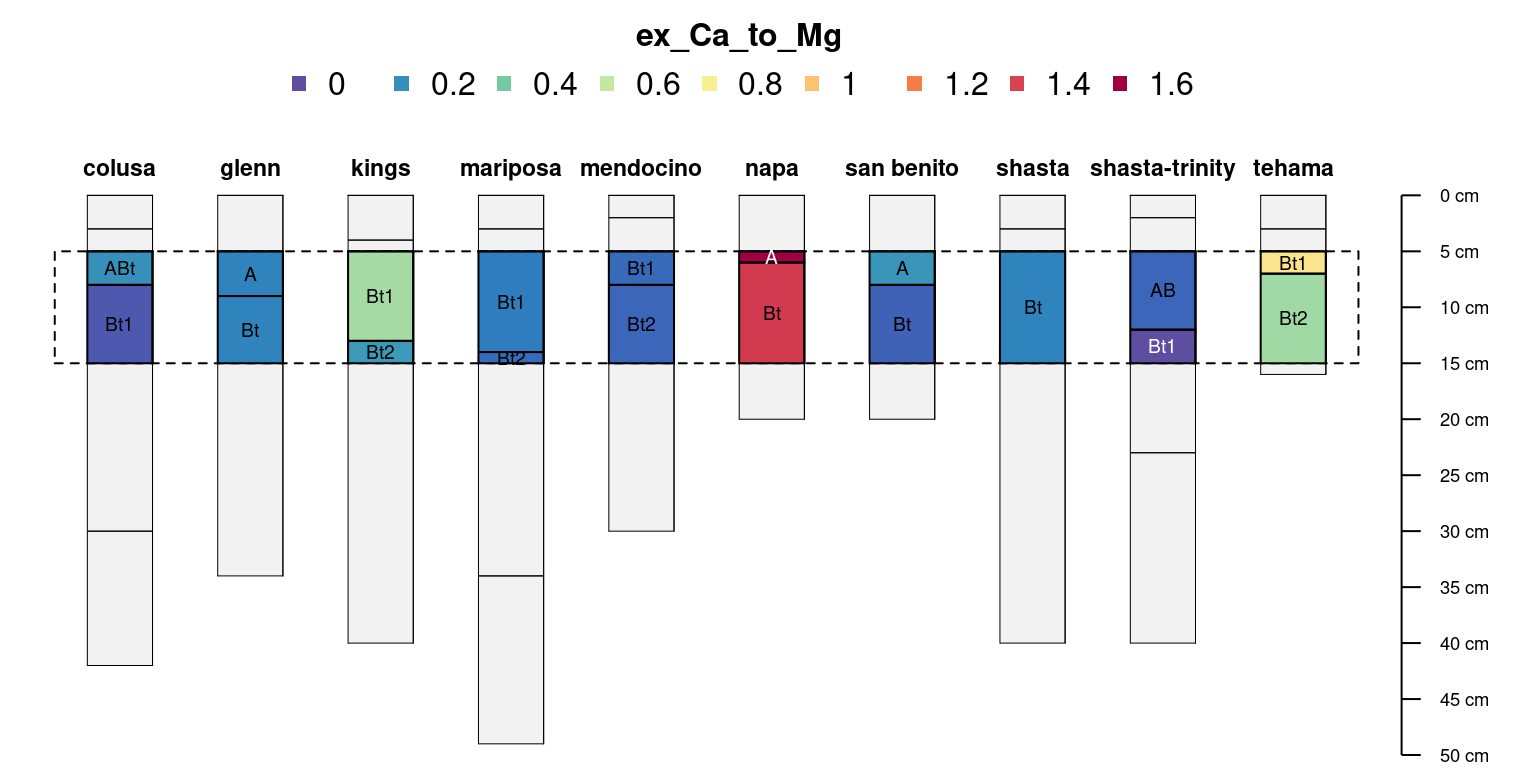
Truncation: trunc()
Truncate the SPC to the interval of 5-15cm:
# truncate to the interval 5-15cm
sp4.truncated <- trunc(sp4, 5, 15)
# plot outlines of original profiles
par(mar = c(0, 0, 3, 1.5))
plotSPC(sp4, color = NA, name = NA, print.id = FALSE, lwd = 0.5)
# overlay truncation criteria
rect(xleft = 0.5, ybottom = 15, xright = length(sp4) + 0.5, ytop = 5, lty = 2)
# add truncated SoilProfileCollection
plotSPC(sp4.truncated, depth.axis = FALSE, add = TRUE, cex.names = 0.6, name = 'name', color = 'ex_Ca_to_Mg', name.style = 'center-center')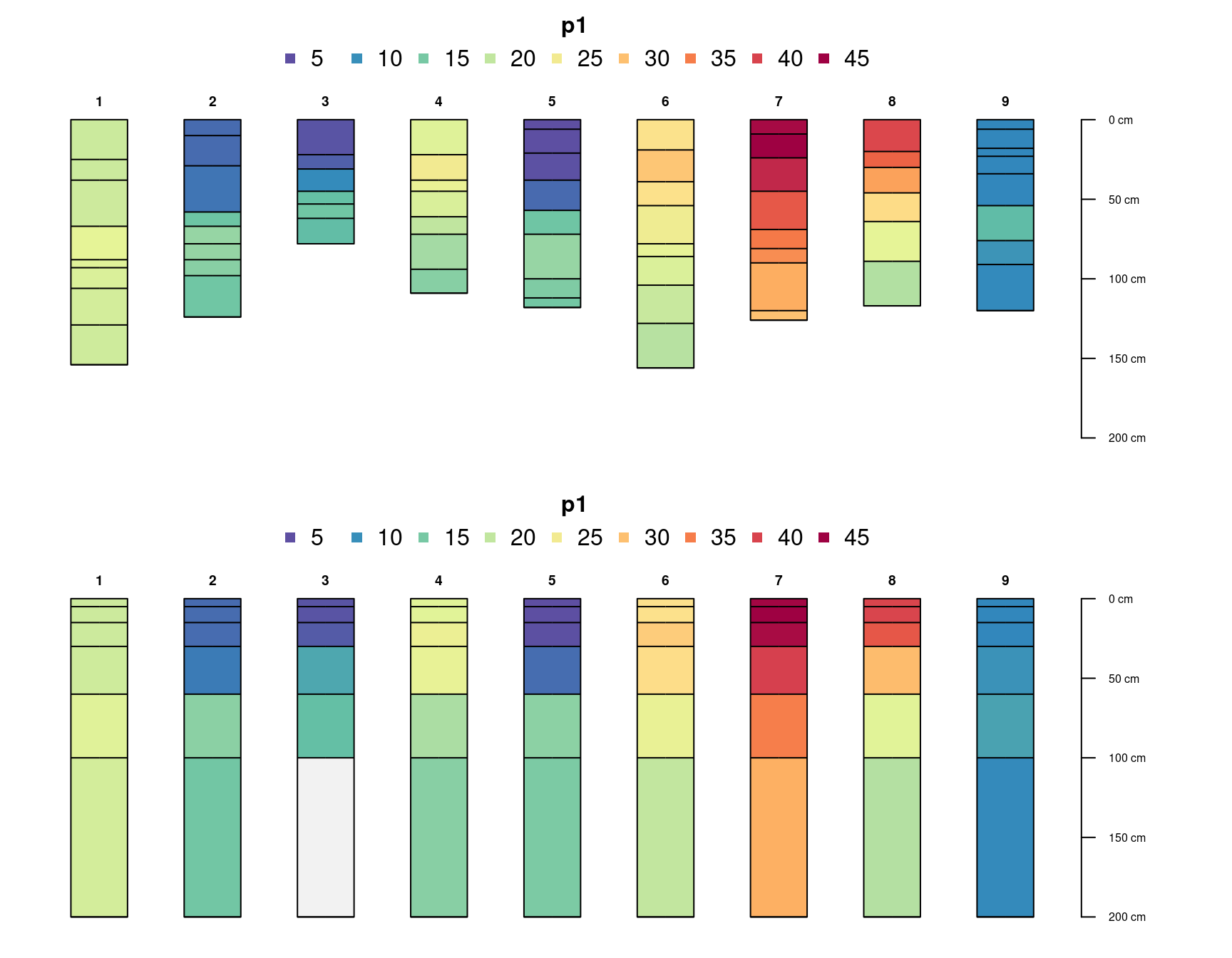
Change of Support
There is often a need for converting data to a new set of depth
intervals. This re-alignment of horizon depths via aggregation can be
considered a type of change of support. The operation on
SoilProfileCollection objects requires some aggregation
function (mean, median, etc.) and possibly interpolation
(e.g. splines).
Aggregation over “slabs”
The slab() function is the simplest way to implement a
change of depth support via aggregation. Profile grouping criteria and
horizon attribute selection is parameterized via formula: either
groups ~ var1 + var2 + var3 where named variables are
aggregated within groups OR where named variables are
aggregated across the entire collection
~ var1 + var2 + var3. The default summary function
(slab.fun) computes the 5th, 25th, 50th, 75th, and 95th
percentiles via Harrell-Davis quantile estimator.
The depth structure (“slabs”) over which summaries are computed is
defined with the slab.structure argument using:
- a single integer (e.g.
10): data are aggregated over a regular sequence of 10-unit thickness slabs - a vector of 2 integers (e.g.
c(50, 60)): data are aggregated over depths spanning 50–60 units - a vector of 3 or more integers
(e.g.
c(0, 5, 10, 50, 100)): data are aggregated over the depths spanning 0–5, 5–10, 10–50, 50–100 units
# aggregate a couple of the horizon-level attributes,
# across the entire collection,
# from 0--10 cm
# computing the mean value ignoring missing data
slab(
sp4,
fm = ~ sand + silt + clay,
slab.structure = c(0, 10),
slab.fun = mean,
na.rm = TRUE
)#> variable all.profiles value contributing_fraction top bottom
#> 1 sand 1 47.63 1 0 10
#> 2 silt 1 31.15 1 0 10
#> 3 clay 1 21.11 1 0 10
# again, this time within groups defined by a site-level attribute:
slab(
sp4,
fm = group ~ sand + silt + clay,
slab.structure = c(0, 10),
slab.fun = mean,
na.rm = TRUE
)#> variable group value contributing_fraction top bottom
#> 1 sand A 48.26 1 0 10
#> 2 sand B 47.00 1 0 10
#> 3 silt A 31.52 1 0 10
#> 4 silt B 30.78 1 0 10
#> 5 clay A 20.30 1 0 10
#> 6 clay B 21.92 1 0 10
# again, this time over several depth ranges
slab(
sp4,
fm = ~ sand + silt + clay,
slab.structure = c(0, 10, 25, 40),
slab.fun = mean,
na.rm = TRUE
)#> variable all.profiles value contributing_fraction top bottom
#> 1 sand 1 47.63000 1.0000000 0 10
#> 2 sand 1 42.38931 0.8733333 10 25
#> 3 sand 1 32.14607 0.5933333 25 40
#> 4 silt 1 31.15000 1.0000000 0 10
#> 5 silt 1 29.41221 0.8733333 10 25
#> 6 silt 1 31.34831 0.5933333 25 40
#> 7 clay 1 21.11000 1.0000000 0 10
#> 8 clay 1 28.10687 0.8733333 10 25
#> 9 clay 1 36.26966 0.5933333 25 40
# again, this time along 1-cm slices, computing quantiles
agg <- slab(sp4, fm = ~ Mg + Ca + ex_Ca_to_Mg + CEC_7 + clay)
# see ?slab for details on the default aggregate function
head(agg)#> variable all.profiles p.q5 p.q25 p.q50 p.q75 p.q95 contributing_fraction top bottom
#> 1 Mg 1 6.015 12.175 14.60 21.125 27.13 1 0 1
#> 2 Mg 1 6.015 12.175 14.60 21.125 27.13 1 1 2
#> 3 Mg 1 6.015 12.175 19.15 25.650 27.76 1 2 3
#> 4 Mg 1 6.195 13.175 21.05 25.050 30.73 1 3 4
#> 5 Mg 1 6.195 13.175 21.05 25.050 30.73 1 4 5
#> 6 Mg 1 6.195 13.175 21.05 26.250 31.72 1 5 6
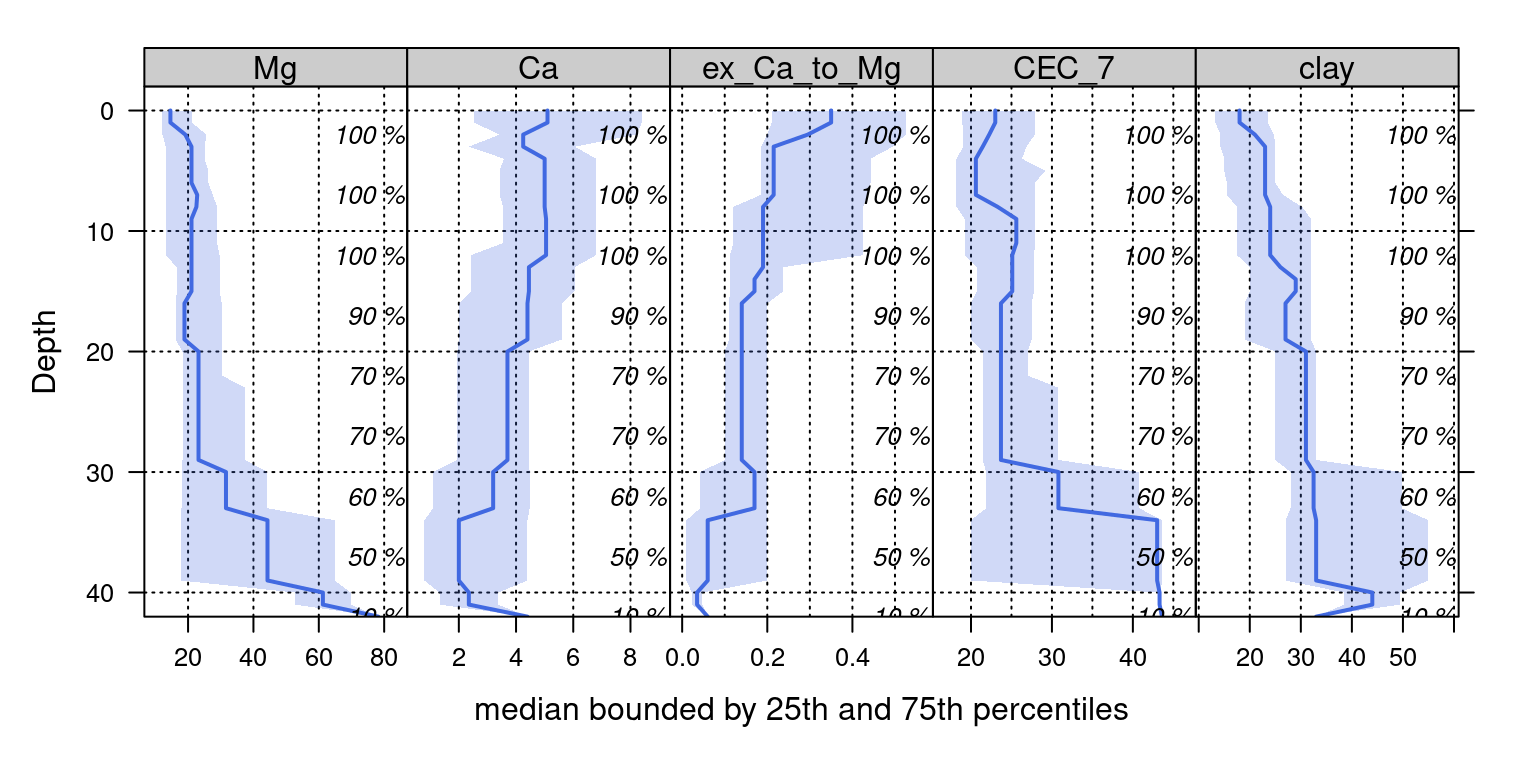
# plot median +/i bounds defined by the 25th and 75th percentiles
# this is lattice graphics, syntax is a little rough
xyplot(top ~ p.q50 | variable, data = agg, ylab = 'Depth',
xlab = 'median bounded by 25th and 75th percentiles',
lower = agg$p.q25, upper = agg$p.q75, ylim = c(42, -2),
panel = panel.depth_function,
alpha = 0.25, sync.colors = TRUE,
par.settings = list(superpose.line = list(col = 'RoyalBlue', lwd = 2)),
prepanel = prepanel.depth_function,
cf = agg$contributing_fraction, cf.col = 'black', cf.interval = 5,
layout = c(5, 1), strip = strip.custom(bg = grey(0.8)),
scales = list(x = list(
tick.number = 4,
alternating = 3,
relation = 'free'
))
)
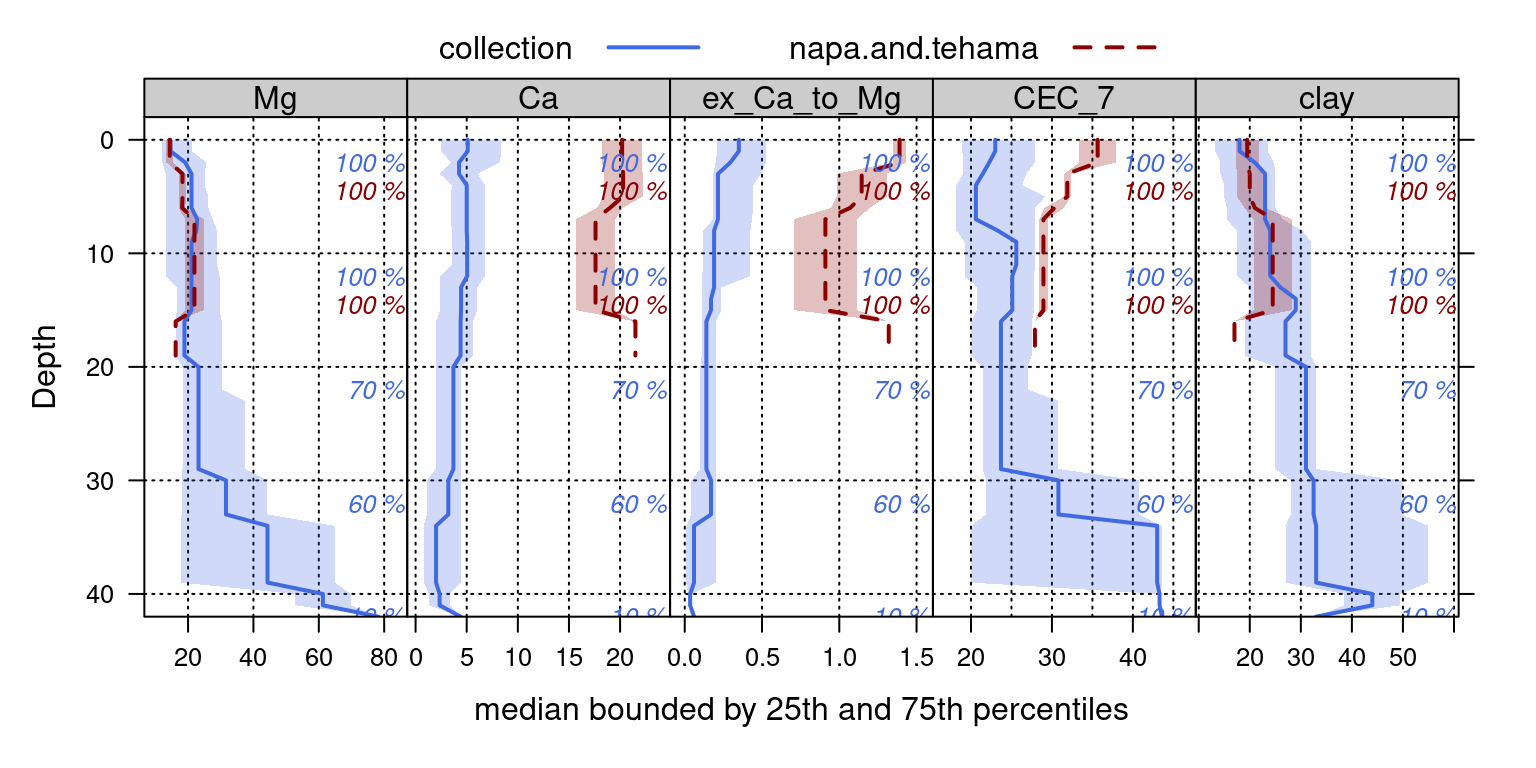
Depth-wise aggregation can be useful for visual evaluation of multivariate similarity among groups of profiles.
# processing the "napa" and tehama profiles
idx <- which(profile_id(sp4) %in% c('napa', 'tehama'))
napa.and.tehama <- slab(sp4[idx,], fm = ~ Mg + Ca + ex_Ca_to_Mg + CEC_7 + clay)
# combine with the collection-wide aggregate data
g <- make.groups(collection = agg, napa.and.tehama = napa.and.tehama)
# compare graphically:
xyplot(top ~ p.q50 | variable, groups = which, data = g, ylab = 'Depth',
xlab = 'median bounded by 25th and 75th percentiles',
lower = g$p.q25, upper = g$p.q75, ylim = c(42, -2),
panel = panel.depth_function,
alpha = 0.25, sync.colors = TRUE, cf = g$contributing_fraction, cf.interval = 10,
par.settings = list(superpose.line = list(
col = c('RoyalBlue', 'Red4'),
lwd = 2,
lty = c(1, 2)
)),
prepanel = prepanel.depth_function,
layout = c(5, 1),
strip = strip.custom(bg = grey(0.8)),
scales = list(x = list(
tick.number = 4,
alternating = 3,
relation = 'free'
)),
auto.key = list(columns = 2,
lines = TRUE,
points = FALSE)
)
The following example is based on a set of 9 randomly generated profiles, re-aligned to the Global Soil Map (GSM) standard depths.
library(data.table)
# 9 random profiles
# 1 simulated property via logistic power peak (LPP) function
# 6, 7, or 8 horizons per profile
# result is a list of single-profile SPC
d <- lapply(
as.character(1:9),
random_profile,
n = c(6, 7, 8),
n_prop = 1,
method = 'LPP',
SPC = TRUE
)
# combine into single SPC
d <- combine(d)
# GSM depths
gsm.depths <- c(0, 5, 15, 30, 60, 100, 200)
# aggregate using mean: wt.mean within slabs
# see ?slab for ideas on how to parameterize slab.fun
d.gsm <- slab(d, fm = id ~ p1, slab.structure = gsm.depths, slab.fun = mean, na.rm = TRUE)
# note: result is in long-format
# note: horizon names are lost due to aggregation
head(d.gsm, 7)#> variable id value contributing_fraction top bottom
#> 1 p1 1 45.44919 1.0000000 0 5
#> 2 p1 1 52.60948 1.0000000 5 15
#> 3 p1 1 50.62579 1.0000000 15 30
#> 4 p1 1 46.70293 1.0000000 30 60
#> 5 p1 1 37.54221 1.0000000 60 100
#> 6 p1 1 31.23363 0.7826087 100 200
#> 7 p1 2 16.26688 1.0000000 0 5A simple graphical comparison of the original and re-aligned soil
profile data, after converting slab() result from long
-> wide format with {data.table} dcast():
# reshape to wide format
# this scales to > 1 aggregated variables
d.gsm.pedons <- data.table::dcast(
data.table(d.gsm),
formula = id + top + bottom ~ variable,
value.var = 'value'
)
# init SPC
depths(d.gsm.pedons) <- id ~ top + bottom
# iterate over aggregated profiles and make new hz names
d.gsm.pedons$hzname <- profileApply(d.gsm.pedons, function(i) {
paste0('GSM-', 1:nrow(i))
})
# compare original and aggregated
par(mar = c(1, 0, 3, 3), mfrow = c(2, 1))
plotSPC(d, color = 'p1')
mtext('original depths', side = 2, line = -1.5)
plotSPC(d.gsm.pedons, color = 'p1')
mtext('GSM depths', side = 2, line = -1.5)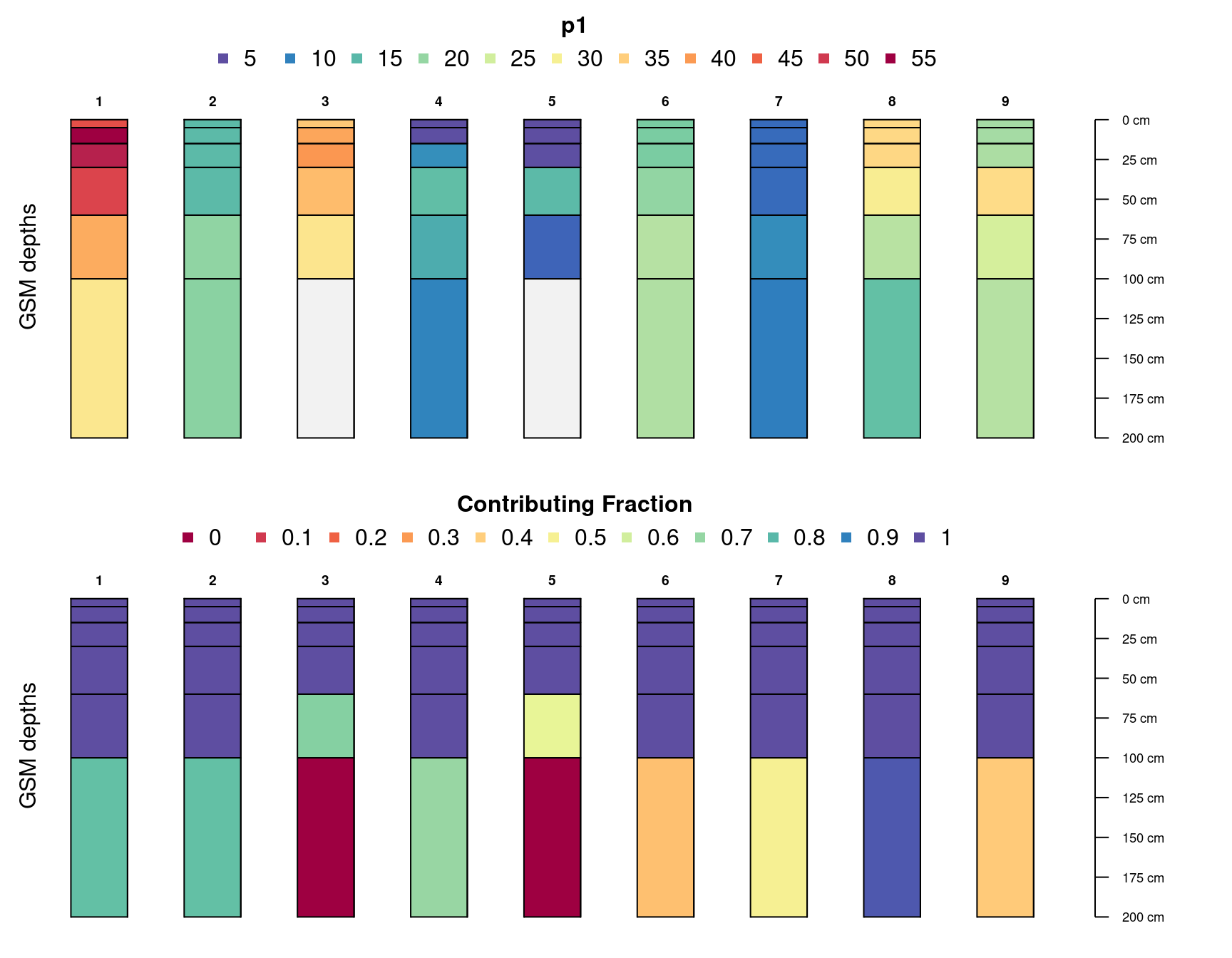
Note that re-aligned data may not represent reality (and should
therefore be used with caution) when the original soil depth is
shallower than the deepest of the new (re-aligned) horizon depths. The
contributing_fraction metric returned by
slab() can be useful for assessing how much real
data were used to generate the new set of re-aligned data.
# reshape to wide format
d.gsm.pedons.2 <- data.table::dcast(
data.table(d.gsm),
formula = id + top + bottom ~ variable,
value.var = 'contributing_fraction'
)
# init SPC
depths(d.gsm.pedons.2) <- id ~ top + bottom
# compare original and aggregated
par(mar = c(1, 0, 3, 3), mfrow = c(2, 1))
plotSPC(d.gsm.pedons, name = '', color = 'p1')
mtext('GSM depths', side = 2, line = -1.5)
plotSPC(
d.gsm.pedons.2,
name = '',
color = 'p1',
col.label = 'Contributing Fraction',
)
mtext('GSM depths', side = 2, line = -1.5)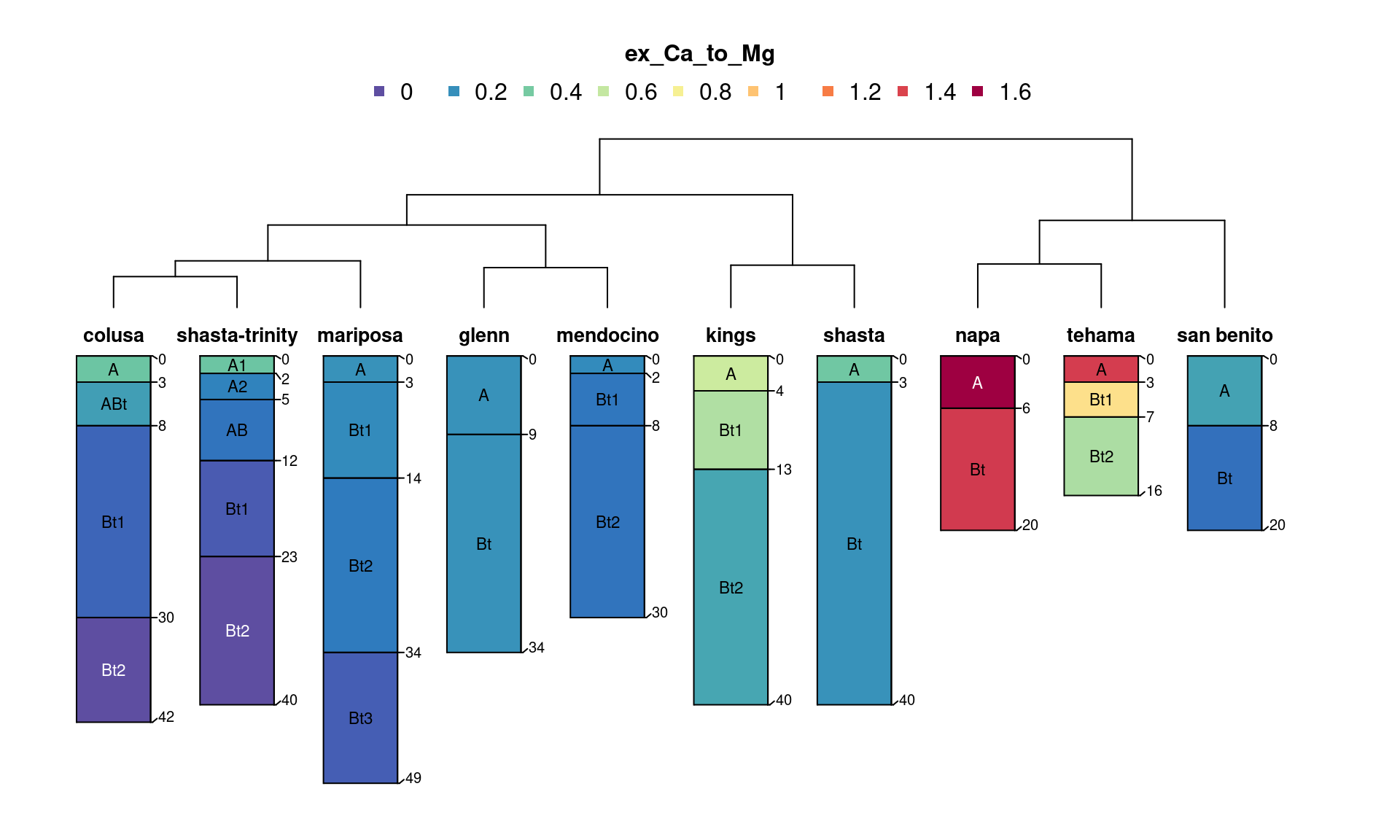
Pair-Wise Dissimilarity: NCSP()
Calculation of between-profile dissimilarity is performed using
NCSP() (Numerical Classification of Soil Profiles). As of
aqp 2.0, NCSP() replaces the older implementation in
profile_compare() which is now deprecated. Dissimilarity
values depend on attributes selection (e.g. clay, CEC, pH , etc.),
optional depth-weighting parameter (k), and a maximum depth
of evaluation (maxDepth).
See the function manual page and this paper for details.
library(cluster)
library(ape)
# start fresh
data(sp4)
depths(sp4) <- id ~ top + bottom
hzdesgnname(sp4) <- 'name'
# eval dissimilarity:
# using Ex-Ca:Mg ratio and CEC at pH 7
# no depth-weighting (k = 0)
# to a maximum depth of 40 cm
d <- NCSP(sp4, vars = c('ex_Ca_to_Mg', 'CEC_7'), k = 0, maxDepth = 40)
# check distance matrix:
round(d, 1)#> colusa glenn kings mariposa mendocino napa san benito shasta shasta-trinity
#> glenn 13.8
#> kings 16.0 13.1
#> mariposa 8.4 11.6 16.5
#> mendocino 12.1 8.2 17.0 15.5
#> napa 31.5 24.9 30.5 30.3 22.2
#> san benito 26.8 21.4 27.4 29.3 16.4 18.0
#> shasta 17.2 13.6 8.7 17.6 17.6 34.8 23.3
#> shasta-trinity 6.4 16.9 22.3 9.6 17.0 30.9 28.3 23.3
#> tehama 30.1 23.9 29.2 28.6 20.8 9.0 15.3 32.7 29.2
# visualize dissimilarity matrix via divisive hierarchical clustering
d.diana <- diana(d)
# may require some margin adjustments
par(mar = c(0, 0, 4, 0))
plotProfileDendrogram(
sp4,
d.diana,
scaling.factor = 0.9,
y.offset = 5,
cex.names = 0.7,
width = 0.3,
color = 'ex_Ca_to_Mg',
name.style = 'center-center',
hz.depths = TRUE,
depth.axis = FALSE
)
Some additional examples can be found in:
This document is based on aqp version 2.3.2.