Annotate diagnostic features within a sketch of soil profiles.
Usage
addDiagnosticBracket(
s,
kind,
feature = "featkind",
top = "featdept",
bottom = "featdepb",
...
)Arguments
- s
SoilProfileCollectionobject- kind
character, filter applied to
featurecolumn of diagnostic horizons registered withins- feature
column name containing feature kind
- top
column name containing feature top depth
- bottom
column name containing feature top depth
- ...
additional arguments passed to
addBracket()
Details
Additional examples can be found in this tutorial.
Note
This is a low-level plotting function: you must first plot a SoilProfileCollection object before using this function.
Examples
# example data
x <- c(
'P1:AAA|BwBwBwBw|CCCCCCC|CdCdCdCd',
'P2:Ap|AA|E|BhsBhs|Bw1Bw1|CCCCC',
'P3:A|Bt1Bt1Bt1|Bt2Bt2Bt2|Bt3|Cr|RRRRR',
'P4:AA|EEE|BhsBhsBhsBhs|BwBw|CCCCC',
'P5:AAAA|ACACACACAC|CCCCCCCCCCC|CdCdCd'
)
s <- quickSPC(x)
diagnostic_hz(s) <- data.frame(
id = c('P1', 'P4'),
t = c(12, 25),
b = c(70, 100),
kind = c('Best', 'Best')
)
op <- par(no.readonly = TRUE)
par(mar = c(0, 0, 3, 2))
# sketches
plotSPC(
s, name = 'name', name.style = 'center-center', cex.names = 0.75, max.depth = 210
)
# note that custom top/bottom depths must be supplied
addDiagnosticBracket(
s, feature = 'kind', kind = 'Best', top = 't', bottom = 'b',
labcol = 'kind',
offset = -0.35, col = 'firebrick', tick.length = 0.04, lwd = 2
)
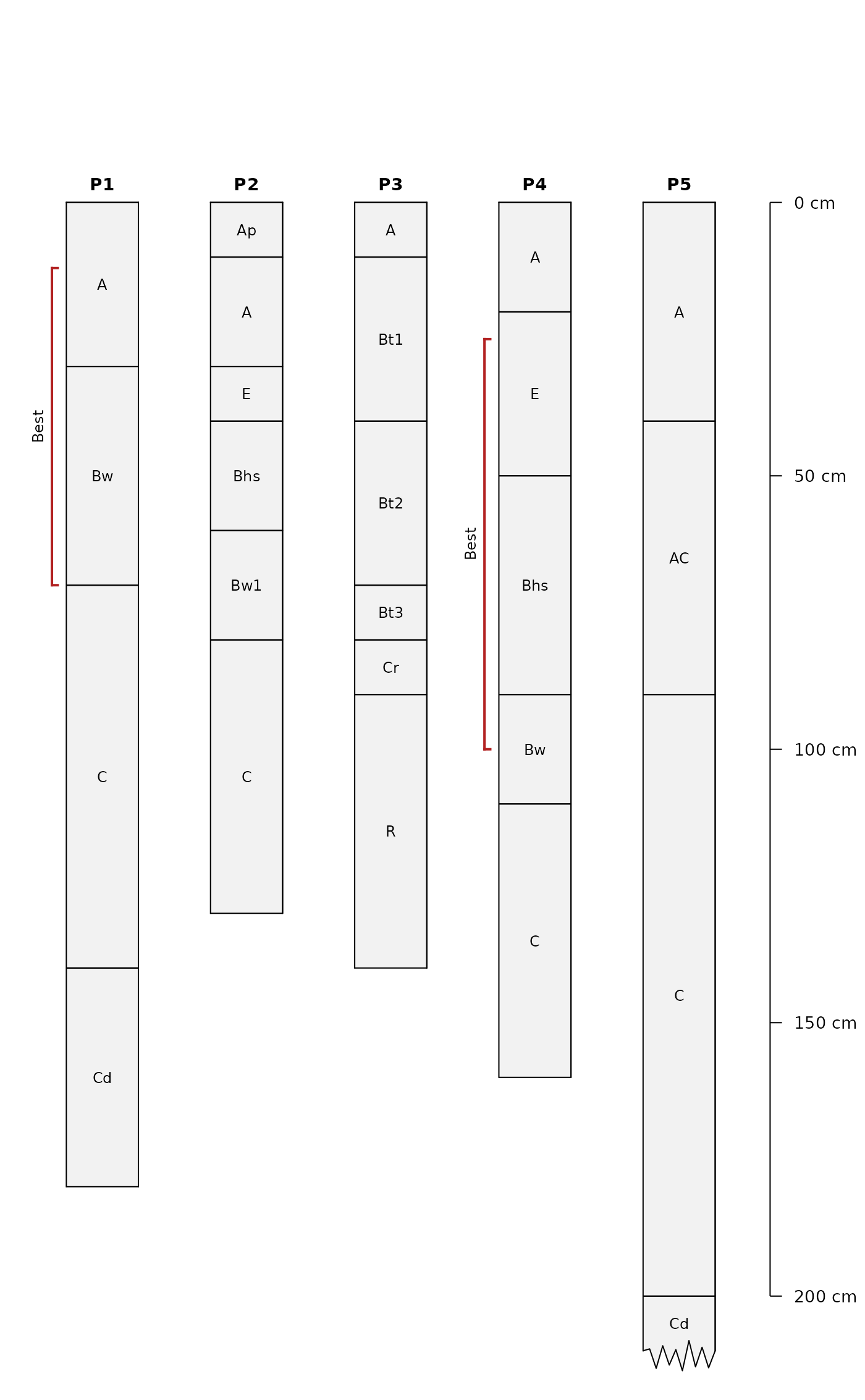 par(op)
par(op)