Generate a discretized vector of genetic horizons along a user-defined
pattern. Deprecated, see dice().
This function is used internally by several higher-level components of the
aqp package. Basic error checking is performed to make sure that
bottom and top horizon boundaries make sense. Note that the horizons should
be sorted according to depth before using this function. The
max_depth argument is used to specify the maximum depth of profiles
within a collection, so that data from any profile shallower than this depth
is padded with NA.
unroll(top, bottom, prop, max_depth, bottom_padding_value = NA, strict = FALSE)Arguments
- top
vector of upper horizon boundaries, must be an integer
- bottom
vector of lower horizon boundaries, must be an integer
- prop
vector of some property to be "unrolled" over a regular sequence
- max_depth
maximum depth to which missing data is padded with NA
- bottom_padding_value
value to use when padding missing data
- strict
should horizons be strictly checked for self-consistency? defaults to FALSE
Value
a vector of "unrolled" property values
References
https://casoilresource.lawr.ucdavis.edu/
Examples
data(sp1)
# subset a single soil profile:
sp1.1 <- subset(sp1, subset=id == 'P001')
# demonstrate how this function works
x <- with(sp1.1, unroll(top, bottom, prop, max_depth=50))
#> Warning: 'unroll' is deprecated.
#> Use 'dice' instead.
#> See help("Deprecated")
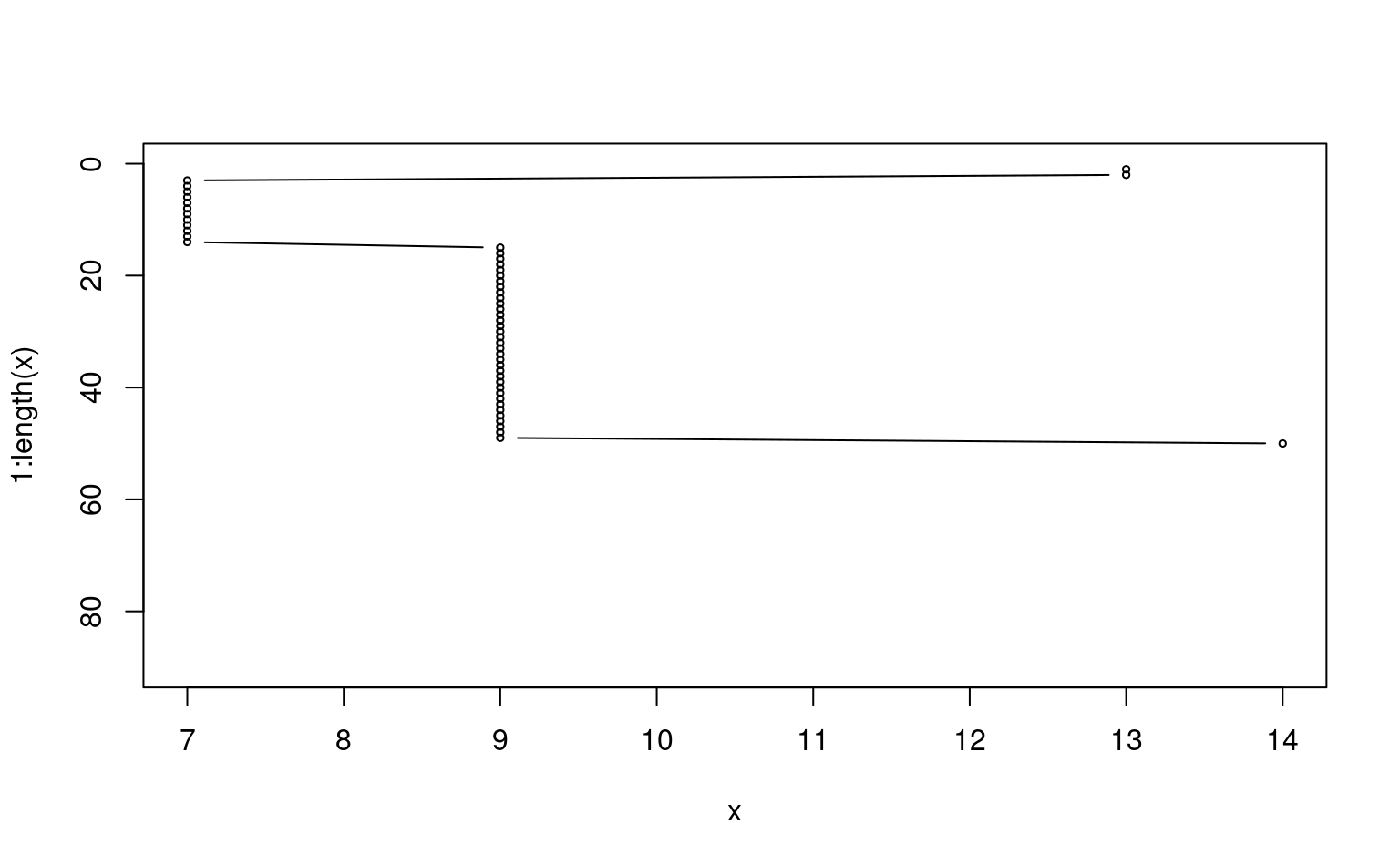
plot(x, 1:length(x), ylim=c(90,0), type='b', cex=0.5)